Press Releases
CEPI-backed Research Preparedness Program West Africa aims to bolster regional clinical trial capacity and disease outbreak readiness. Partnership will support regional stakeholders to strengthen clinical research capacity to conduct Phase 2b / 3 clinical trials of Lassa fever vaccines in the region, for the region; […]
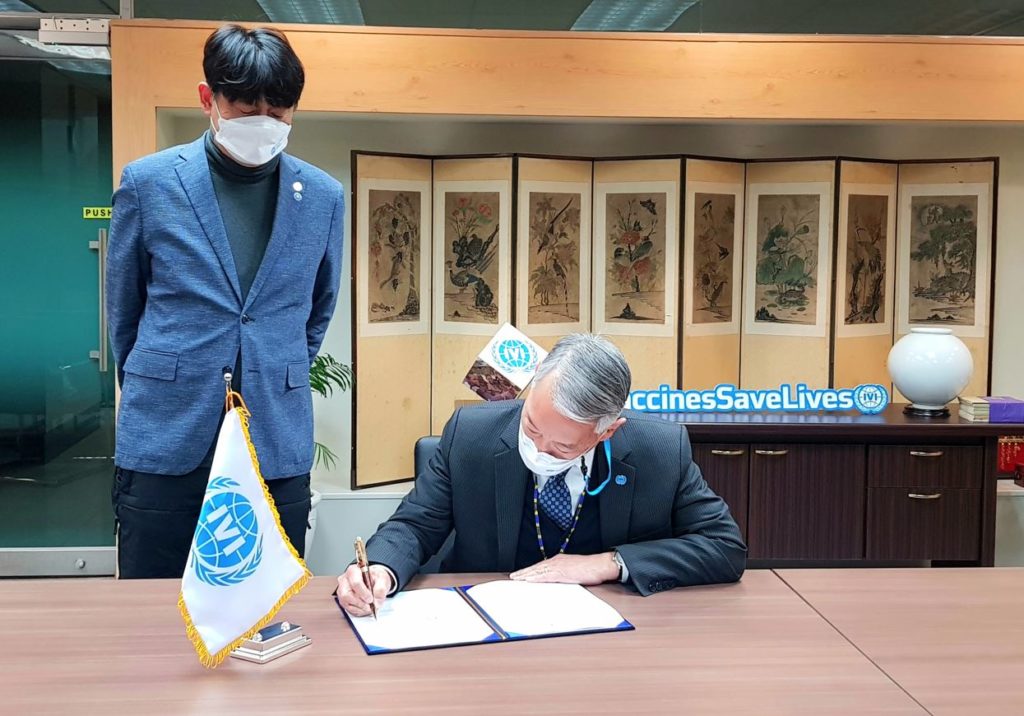
Joining forces in four main areas including vaccine immune response analysis using KAIST’s technology The International Vaccine Institute (IVI) and the Korea Advanced Institute of Science and Technology (KAIST) exchanged a memorandum of understanding (MOU) for global vaccine research collaboration at KAIST headquarters in […]
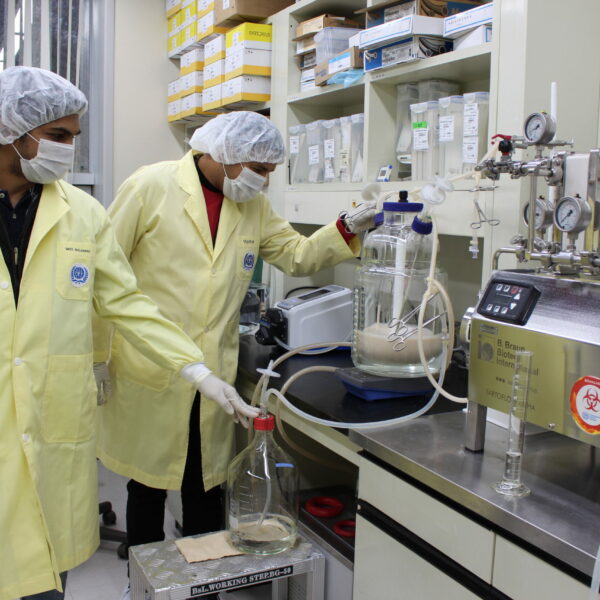
IVI held an opening ceremony at headquarters in Seoul on October 30, 2023, to kickstart Hands-on Training for Upstream Process in Vaccine Manufacturing in partnership with KEMRI-Wellcome Trust Research Programme (KWTRP) and Korea Biopharmaceutical CMO (K-Bio CMO). Credit: IVI October 30, 2023 – SEOUL, […]
September 14, 2023, HONG KONG SAR and SEOUL, Republic of Korea – The International Vaccine Institute (IVI), an international organization with a mission to discover, develop, and deliver safe, effective, and affordable vaccines for global health, and Grid Biosciences (Grid), a biotech company developing […]
June 21, 2023, SEOUL, Republic of Korea – The International Vaccine Institute (IVI) and Seoul National University signed an agreement to expand collaboration in research. IVI is the only international organization dedicated to the development and delivery of new vaccines to protect people […]
Report contains key findings for stakeholders engaging in AMR surveillance, research, policy, regulatory decision-making, and other infectious disease prevention and control programs in Bangladesh and wider Asia, supported by the Fleming Fund. June 20, 2023, SEOUL, Republic of Korea – CAPTURA is a […]
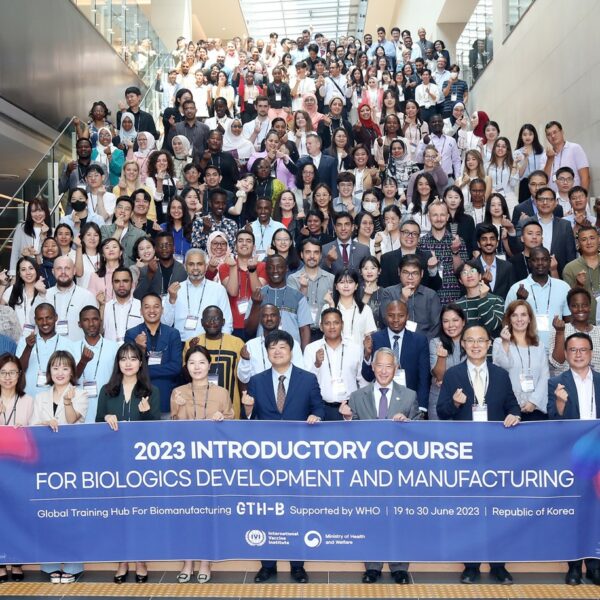
2-week training focuses on technology transfer, talent development, facility management, and aseptic processing Part of the Korean MOHW’s Global Training Hub for Biomanufacturing, the program aims to strengthen local bio production capabilities in LMICs to address vaccine inequity and strengthen pandemic preparedness June […]
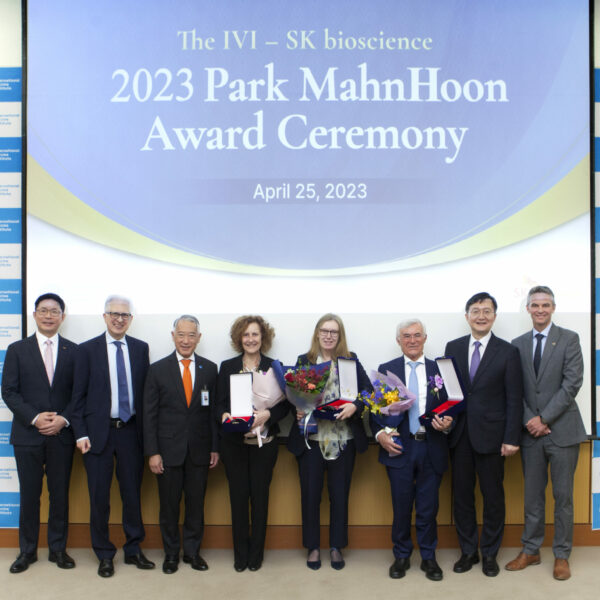
Awardees give lectures to highlight their work and achievements, share knowledge at the Award Forum April 26, 2023 – SEOUL, Korea – The International Vaccine Institute (IVI) gave the 2023 IVI-SK bioscience Park MahnHoon Award to Drs. Rino Rappuoli and Mariagrazia Pizza as […]
Seegene to handle logistics support to test 50,000 girls and women ages 9-50 in Asia and Africa from August Seegene’s Allplex™ HPV28 Detection tests to be used to improve access to HPV screening Project funded by the Bill & Melinda Gates Foundation, conducted jointly by […]
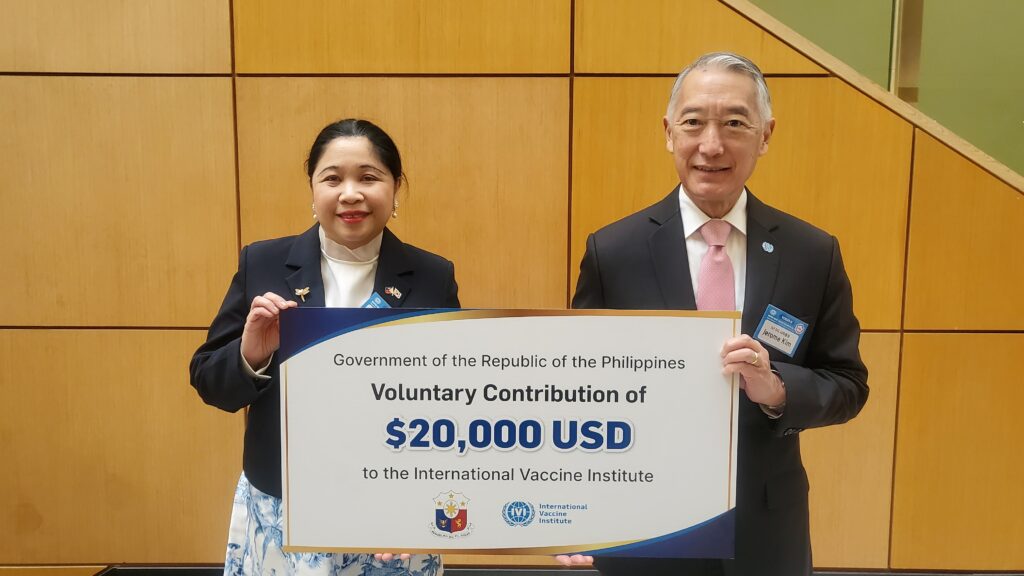
March 14, 2023 – SEOUL, Republic of Korea – The Philippine Ambassador to the Republic of Korea, H.E. Ma. Theresa Dizon-de Vega, announced that the Government of the Republic of the Philippines has made a voluntary contribution to the International Vaccine Institute, an international […]
Provides training for about 40 Afrigen personnel on overall biopharmaceutical production as part of GTH-B program December 13, 2022, SEOUL, Republic of Korea — The International Vaccine Institute (IVI) conducted on-site training and consultation for Afrigen Biologics & Vaccines (Afrigen) in South Africa to help […]
November 10, 2022, SEOUL, Republic of Korea – The International Vaccine Institute (IVI) announced on November 10 it signed a memorandum of understanding (MOU) with Lemonex, a company specializing in RNA gene therapy development, and agreed to seek cooperation in research and development. IVI and Lemonex […]
September 1, 2022, SEOUL, Republic of Korea – The International Vaccine Institute (IVI), an international organization with a mission to discover, develop, and deliver safe, effective, and affordable vaccines for global health, opened the IVI Europe Regional Office (IERO) in Stockholm today. IERO, housed temporarily […]
New evidence supports the use of alternative dosing formulations, expanding TCV delivery options in public health programs June 14, 2022 – SEOUL, Republic of Korea – A new study jointly conducted by the International Vaccine Institute (IVI) and collaborators shows multidose and single-dose […]
May 23, 2022– SEOUL, Republic of Korea and ADDIS ABABA, Ethiopia – The International Vaccine Institute (IVI), an international organization with a mission to discover, develop, and deliver safe, effective, and affordable vaccines, and Armauer Hansen Research Institute (AHRI), established by the Government of […]
Recombinant protein-based ‘SKYCovione™’ adjuvanted with GSK’s pandemic adjuvant demonstrates superior neutralizing titers compared to a control vaccine and safety SK bioscience submits a biologics license application to KMFDS, and will submit license applications to international regulatory agencies Company set to supply 10 million doses of […]
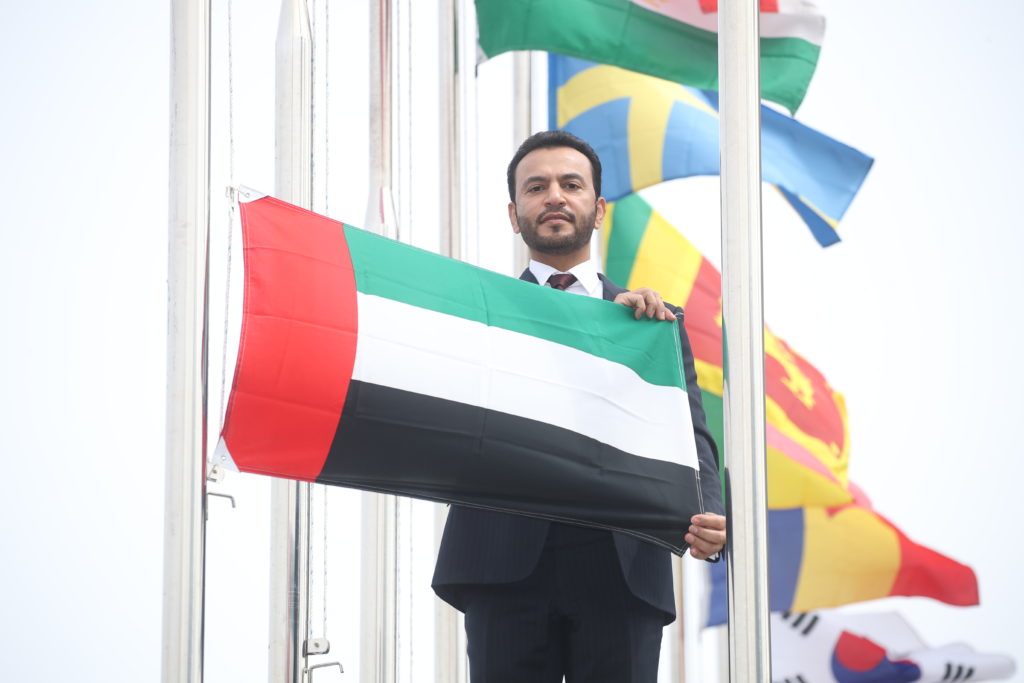
March 17, 2022 – SEOUL, South Korea – The International Vaccine Institute (IVI) announced today that the United Arab Emirates (UAE) joined IVI, becoming the international organization’s 37th member state. His Excellency Abdulla Saif Al Nuaimi, Ambassador of the UAE to the Republic of […]
Institute to run training program on vaccines and biologics development and manufacturing and GxP for 450 trainees from LMICs and Korea February 16, 2022 – SEOUL, Republic of Korea – The International Vaccine Institute (IVI) has been designated by the Ministry of Health and Welfare […]
January 27, 2022 – SEOUL, Republic of Korea – A new study shows that late booster dosing with Vi polysaccharide conjugated with diphtheria toxoid (Vi-DT), one of the typhoid conjugate vaccines (TCVs), at 27.5 months post-first dose is safe and elicits robust immune responses in […]
IVI and the National Vaccine Institute (NVI) of Thailand signed a Definitive Agreement to strengthen their collaborative partnership January 20, 2022 – SEOUL, Republic of Korea – The International Vaccine Institute (IVI) and the National Vaccine Institute (NVI) of Thailand signed a Definitive […]
Recruitment of global Phase III clinical trial of ‘GBP510’ completed, with vaccine expected to be authorized in the first half of 2022 A booster dose study is concurrently being planned to bring additional vaccine to the global COVID-19 vaccine market January 19, 2022 SEOUL, […]
Study will assess the safety and immunogenicity of a heterologous regimen of two approved COVID-19 vaccines in adults Aims to mitigate vaccine shortages and support flexible immunization programs December 20, 2021, SEOUL, Republic of Korea; MAPUTO, Mozambique; OSLO, Norway – The International Vaccine Institute […]
IVI and the Government Offices of Sweden signed a Memorandum of Understanding today to establish the office in Stockholm, creating a European hub for global health research and innovation December 17, 2021, SEOUL, Republic of Korea – The International Vaccine Institute (IVI), an international […]
The webinar “Curbing the Invisible Pandemic: Effective Solutions to Collectively Combat Antimicrobial Resistance” was held on December 7, 2021 December 7, 2021 — The International Vaccine Institute (IVI), Asian Development Bank Southeast Asia Development Solutions (ADB SEADS), Institut Pasteur Korea (IPK), the […]
Cited for contributions to development of vaccines, joining 8 foreign residents to receive Honorary Citizenship in 2021 Second Honorary Seoul Citizen from IVI SEOUL, Korea — Dr. Sushant Sahastrabuddhe, Associate Director General at the International Vaccine Institute (IVI), has been awarded “Honorary Citizenship of […]
IVI, SK sign an agreement to launch the ‘Park MahnHoon Award’ to honor individuals and organizations that made contributions to the vaccine industry Award Ceremony will be held annually in April; IVI and SK ‘committed to establish the new award as a prestigious award for […]
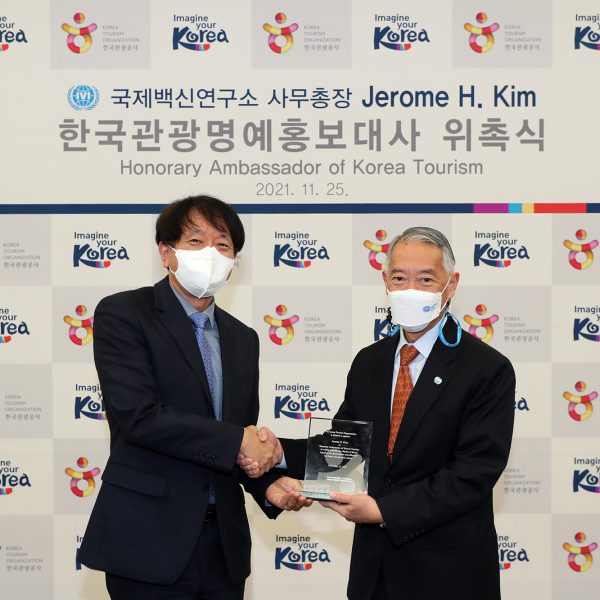
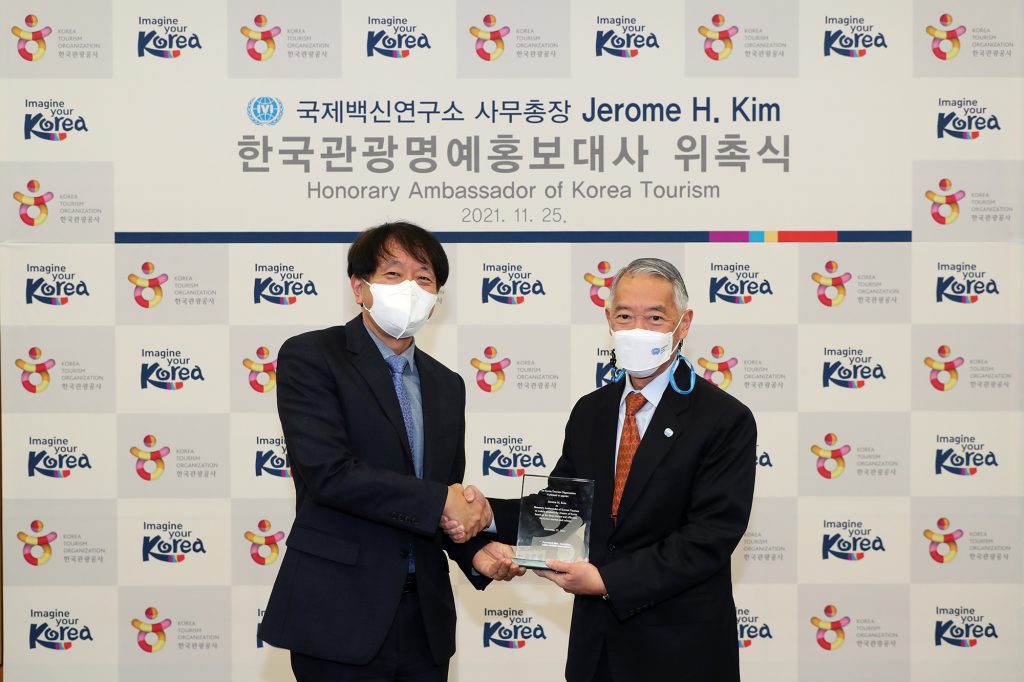
To promote the safety and attractiveness of traveling Korea amidst gradual restart of global tourism The Ministry of Culture, Sports and Tourism (Minister Hwang Hee) and the Korea Tourism Organization (KTO, President Ahn Young-bae) appointed Dr. Jerome H. Kim, Director General of the […]
The International Vaccine Institute (IVI) has been honored with the Presidential Citation of the Republic of Korea’s Development Cooperation Award, in recognition of IVI’s contributions to global health and Korea’s official development assistance. IVI has been cited as being an “International organization focused […]
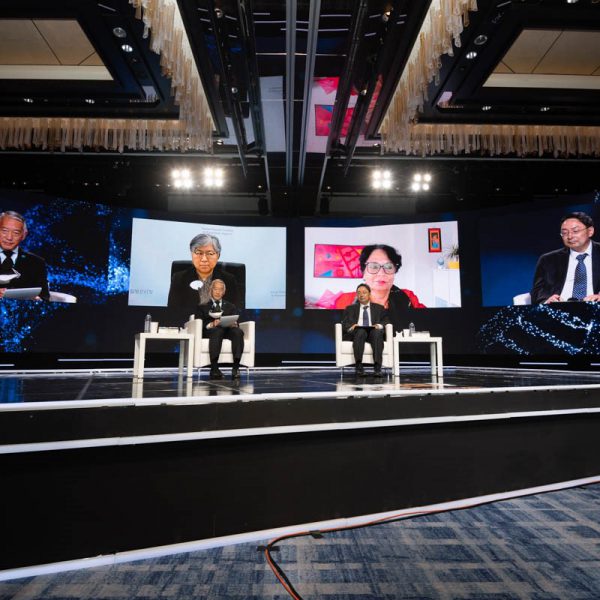
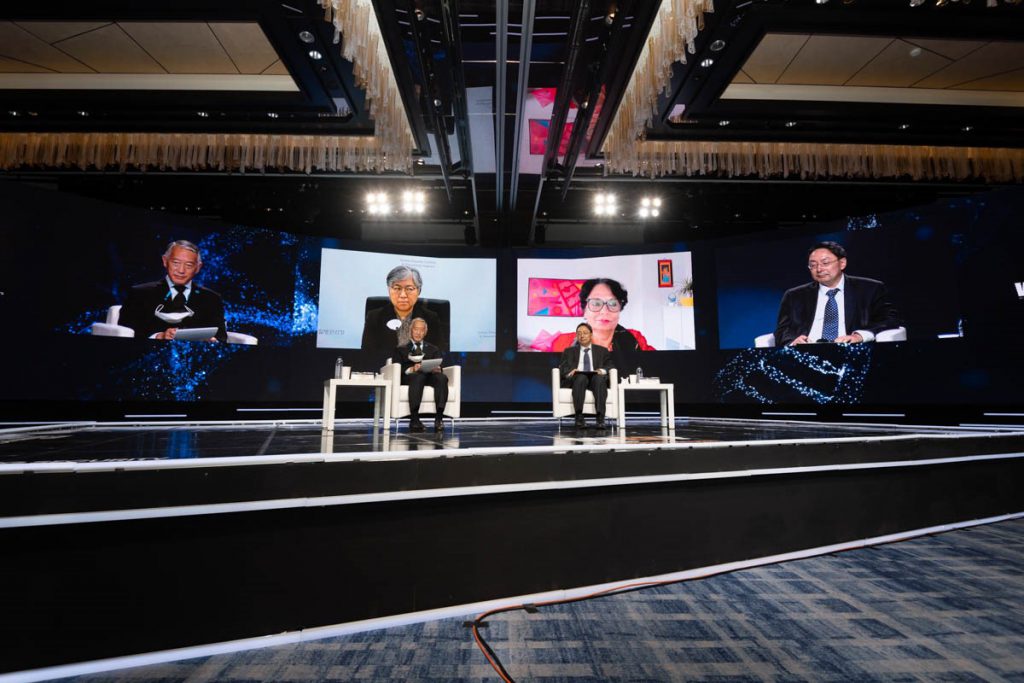
Dr. Jerome Kim, Director General and Dr. Song Manki, Deputy Director General of Science of the International Vaccine Institute (IVI) joined distinguished global leaders and experts as moderator and speaker at the first World Emerging Security Forum, which took place in Seoul on November […]
October 25, 2021 – SEOUL, Republic of Korea, PARIS, France, BRUSSELS, Belgium, BERLIN, Germany – The International Vaccine Institute (IVI) and the Joint European Disruptive Initiative (JEDI) signed a Memorandum of Understanding to establish a collaborative relationship. Both organizations are dedicated to advancing innovations in […]
Until Everyone Is Safe: Global Vaccine Needs and IVI’s Capabilities was livestreamed on October 7, 2021 at 16:00 Korea Standard Time, and a recording of the event is available here. October 7, 2021, SEOUL, Republic of Korea — The International Vaccine […]
Sept. 28, 2021 – SEOUL, South Korea, and HOUSTON, Texas, USA – The International Vaccine Institute (IVI), the Texas Children’s Hospital Center for Vaccine Development, and Baylor College of Medicine’s National School of Tropical Medicine signed a Memorandum of Understanding to promote and further academic […]
International Vaccine Institute (IVI) will lead the clinical trial in Nepal to assess the safety, efficacy and immunogenicity of an adjuvanted recombinant-protein COVID-19 vaccine candidate Phase 3 international clinical trial includes volunteers from several countries, including sites in the US, Asia, Africa and Latin America […]
September 10, 2021, SEOUL, Korea — The International Vaccine Institute’s (IVI) 20th International Vaccinology Course (IVC) concluded today, bringing together 7,350 registered trainees and 27 faculty members for 5 days of online lectures on a range of topics related to the science of vaccines with […]
IVI is leading the Global Chikungunya vaccine Clinical Development Program (GCCDP) consortium in partnership with Bharat Biotech International Ltd. and with support from the Coalition for Epidemic Preparedness Innovations (CEPI) and Ind-CEPI. August 24, 2021 – SEOUL, Republic of Korea – The International Vaccine […]
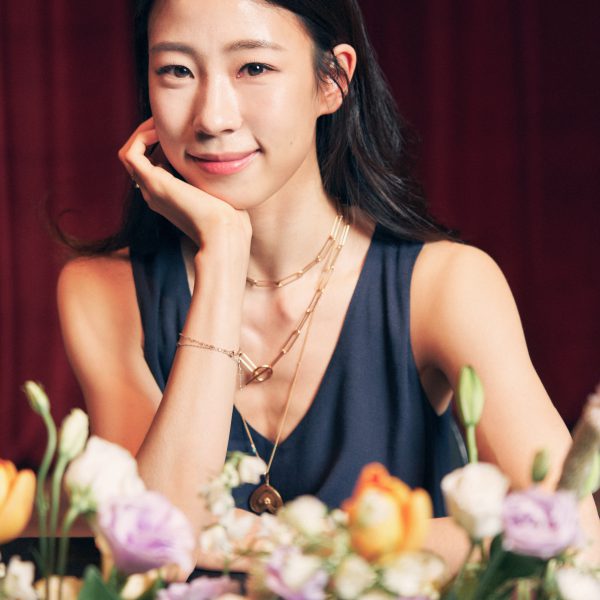
August 18, 2021 – SEOUL, Republic of Korea – The International Vaccine Institute (IVI) named Sae Eun Park, an award-winning ballet dancer from Korea, an IVI Goodwill Ambassador during a virtual ceremony in Seoul today. Ms. Park was named danseuse étoile, the highest-level dancer, […]
July 26, 2021 – SEOUL, Republic of Korea – The International Vaccine Institute (IVI) announced today the start of a cluster-randomized, controlled Phase IV trial to assess the effectiveness of Typbar® typhoid conjugate vaccine (TCV) in preventing typhoid infection in children in Asante Akim, Ghana […]
July 20, 2021; Oslo, Norway and Seoul, Republic of Korea: The Coalition for Epidemic Preparedness Innovations (CEPI) and the International Vaccine Institute (IVI) today announced a new programme of clinical research which aims to expand access to COVID-19 vaccines in Africa. CEPI will provide funding […]
July 15, 2021 – SEOUL, South Korea – The International Vaccine Institute (IVI) hosted a ceremony at headquarters today honoring the People’s Republic of Bangladesh’s ratification of the IVI Establishment Agreement. Over 20 years of vaccine research and capacity-building initiatives with partners in Bangladesh […]
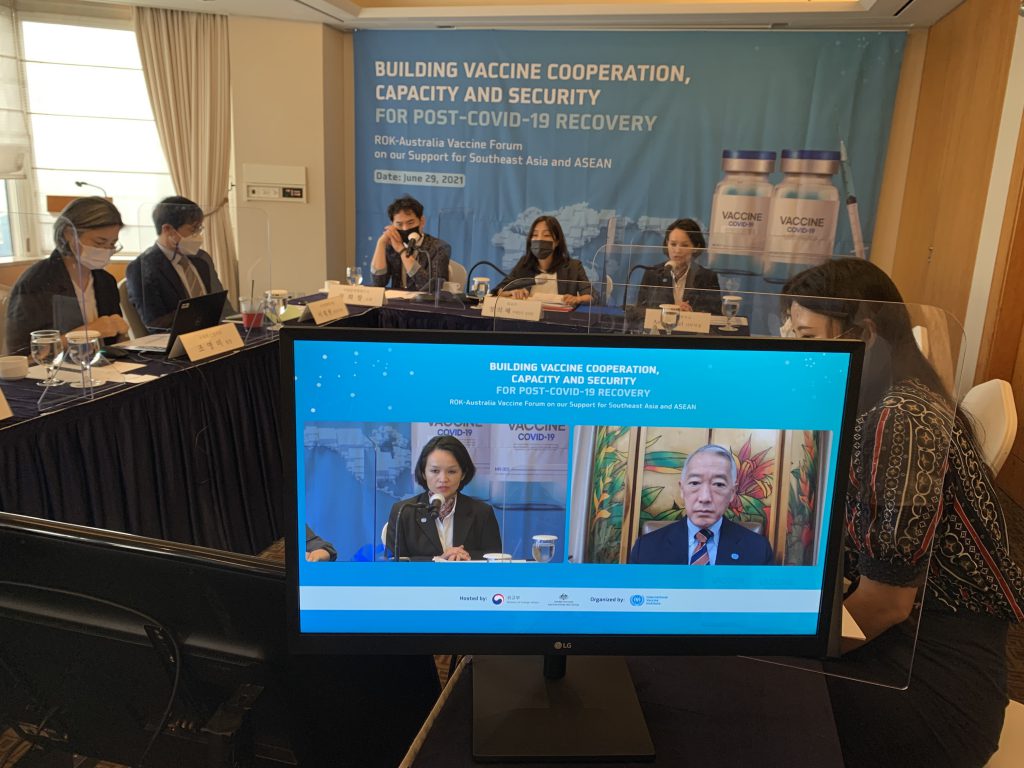
The ROK-Australia-ASEAN Vaccine Forum took place in hybrid live/virtual format with transmission from Seoul on June 29, 2021 June 29, 2021 – SEOUL, Republic of Korea – The International Vaccine Institute (IVI) organized, and the Republic of Korea’s Ministry of Foreign Affairs (MOFA) and […]
May 31, 2021 – SEOUL, South Korea – The International Vaccine Institute (IVI) announced today that the Bill & Melinda Gates Foundation awarded a grant to IVI to develop an adaptive trial design protocol for a Phase 1b/2a clinical trial of a schistosomiasis vaccine. […]
The agreement is an effort to further scientific research in infectious diseases and make knowledge more accessible through advocacy and joint publications. May 19, 2021 – SEOUL, South Korea, RIYADH, Saudi Arabia – The International Vaccine Institute (IVI) and the Future Investment Initiative Institute […]
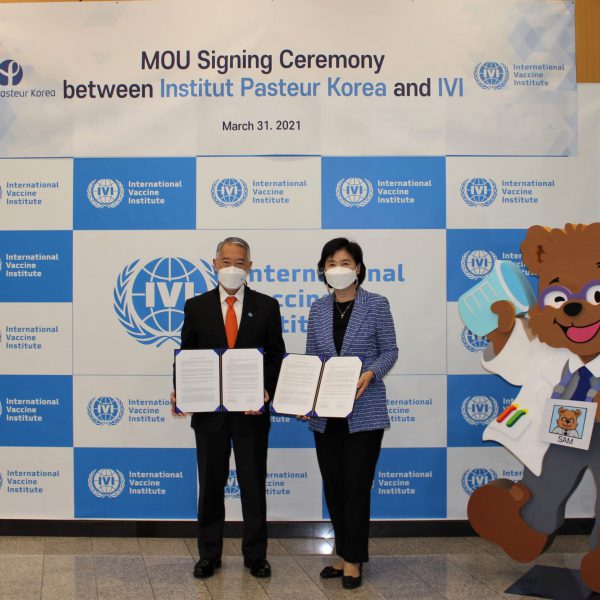
April 1, 2021, GYEONGDO-DO and SEOUL, Korea – The International Vaccine Institute (IVI) and the Institut Pasteur Korea (IPK) signed a memorandum of understanding (MOU) for mutual collaboration in the research and development of therapeutics and vaccines for infectious diseases. The MOU signing ceremony, […]
March 9, 2021 – SEOUL, South Korea – The International Vaccine Institute (IVI) welcomes the Cabinet of Bangladesh’s decision to ratify the IVI Establishment Agreement on February 22, 2021, completing the full accession process to IVI and becoming its 19th State Party. Bangladesh is a founding signatory to IVI’s […]
George Bickerstaff re-elected Chairperson of the International Vaccine Institute’s Board of Trustees
February 22, 2021, SEOUL, Korea – The International Vaccine Institute (IVI) announced today that its Board of Trustees (BOT) re-elected Mr. George Bickerstaff as Chairperson of the BOT. His second three-year term will begin this May. Mr. Bickerstaff has served on the IVI Board […]
IVI to analyze samples from Phase 1/2a trials of Cellid’s COVID-19 vaccine to evaluate immunogenicity January 5, 2021, SEOUL, Korea – Cellid and the International Vaccine Institute (IVI) have exchanged a collaborative research agreement to analyze the immunogenicity of the COVID-19 vaccine “AdCLD-CoV19”. […]
Primary analysis also confirms safety of Vi-DT December 17, 2020 – SEOUL, South Korea – Vi-DT typhoid conjugate vaccine, developed jointly by the International Vaccine Institute (IVI) and SK bioscience, has met the primary endpoints in a phase III study in Nepal. The primary […]
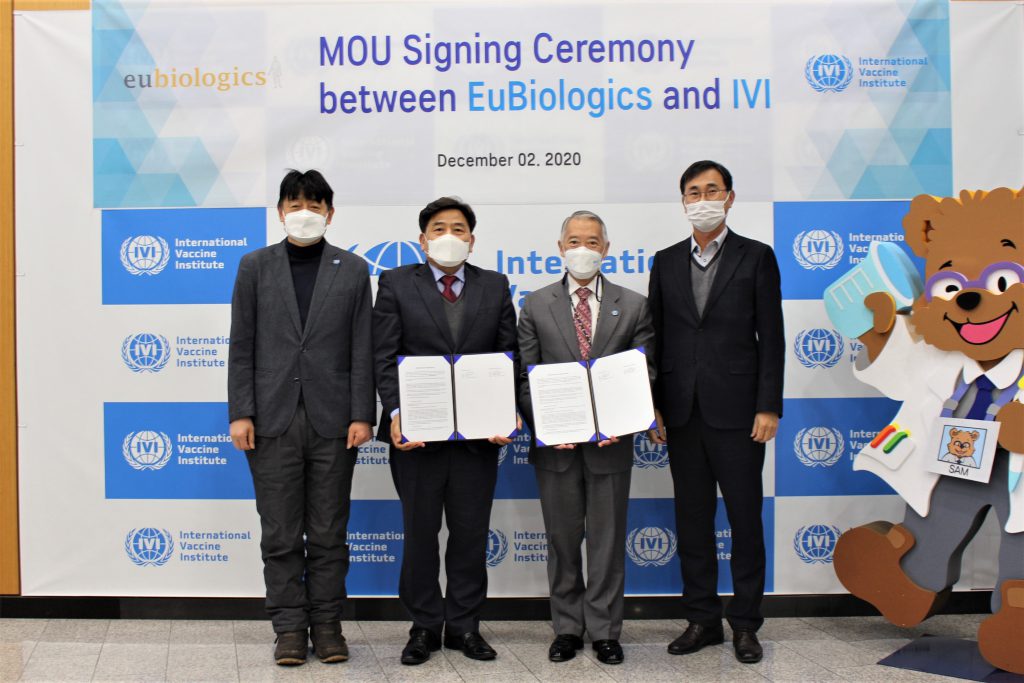
December 3, 2020, SEOUL, South Korea – The International Vaccine Institute (IVI) and EuBiologics exchanged an MOU to cooperate in clinical development of the COVID-19 vaccine the company is currently developing. The signing ceremony at IVI headquarters on December 2 was attended by Dr. Jerome […]
Free webinar, Evidence to Action: Advancing the Antimicrobial Resistance Agenda during a Pandemic, will be held on Thursday, December 3, 2020 at 9:00 Central European Time (17:00 Korean Standard Time). Registration available at ivi.int/evidence-to-action/ December 1, 2020, SEOUL, Korea — The International Vaccine Institute […]
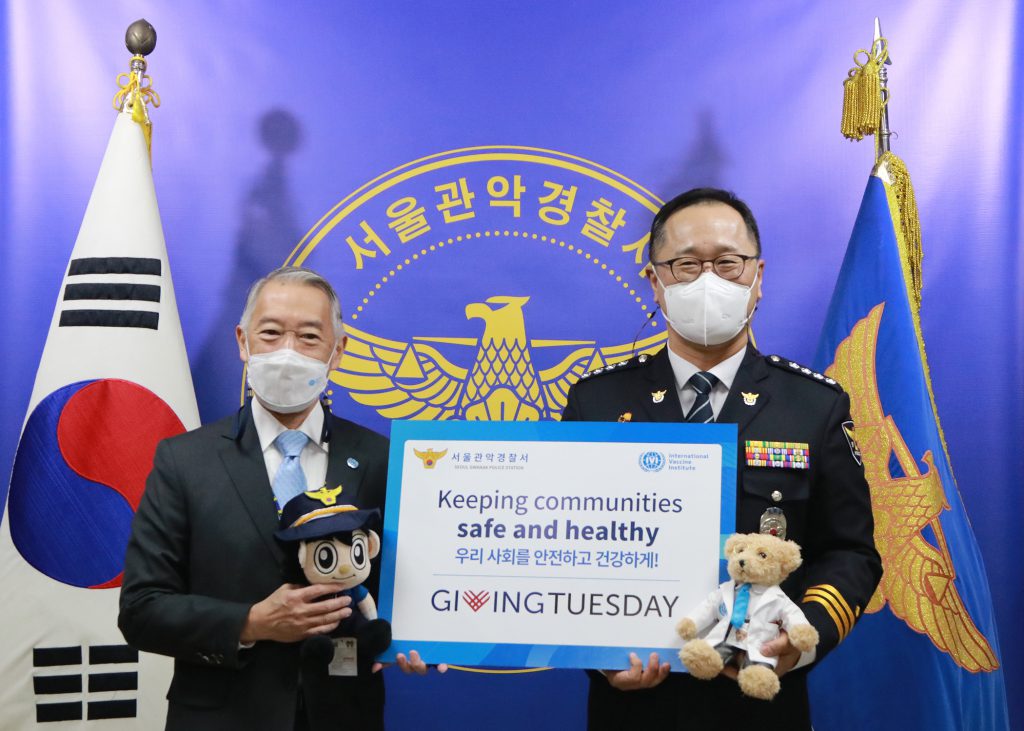
December 1, 2020, SEOUL, South Korea – The International Vaccine Institute (IVI) visited the Seoul Gwanak Police Station today to recognize the Foreign Affairs section for their ongoing support for IVI’s activities and to deliver protective face masks and medical supplies for staff and volunteers. […]
GDEF’s US$8.05 million grant to support ECHO projects to prevent and control cholera and contribute to ‘Ending Cholera—A Global Roadmap to 2030’ The International Vaccine Institute (IVI) and the Republic of Korea’s Global Disease Eradication Fund (GDEF) have agreed to conduct joint projects to […]
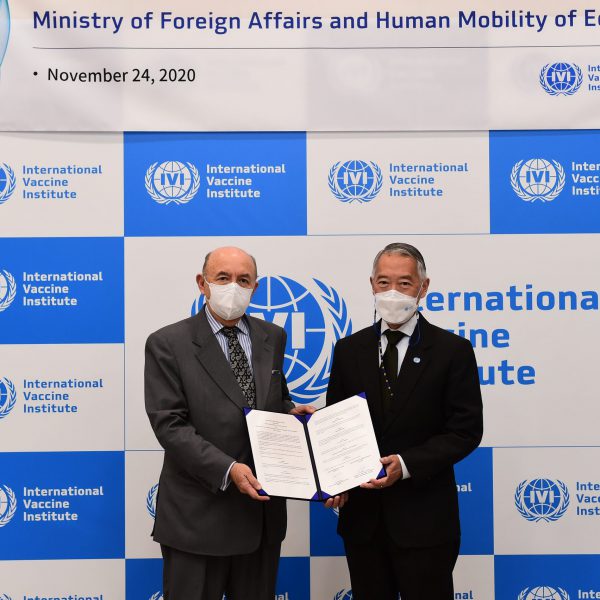
November 24, 2020, SEOUL, Republic of Korea — The International Vaccine Institute (IVI) and the Ministry of Foreign Affairs and Human Mobility of Ecuador exchanged a memorandum of understanding (MOU) today at IVI headquarters in Seoul, Republic of Korea to explore areas of collaboration in […]
Project to provide vaccination for 40,000 residents in areas at risk of cholera and establish disease monitoring system with the Ethiopian Ministry of Health through the Armauer Hansen Research Institute (AHRI) To contribute to health authorities in policymaking for disease prevention by investigating waterborne diseases […]
Aim to accelerate R&D and globalization of Korean vaccines to increase contributions to global health The International Vaccine Institute (IVI) and the Vaccine Innovative Technology ALliance Korea (VITAL-Korea) agreed to join forces to promote vaccine research and development for global health. IVI […]
October 28, 2020, SEOUL, Korea — The International Vaccine Institute (IVI) received an award by the Minister of Health and Welfare of the Republic of Korea for achievements in biosafety management. IVI was honored with the Minister of Health and Welfare Award for achievements in […]
The International Vaccine Institute (IVI) is partnering with the World Sanity Foundation (WSF) in a campaign to encourage the use of face masks in an effort to reduce the spread of COVID-19 cases until a safe and effective vaccine becomes widely available. The first […]
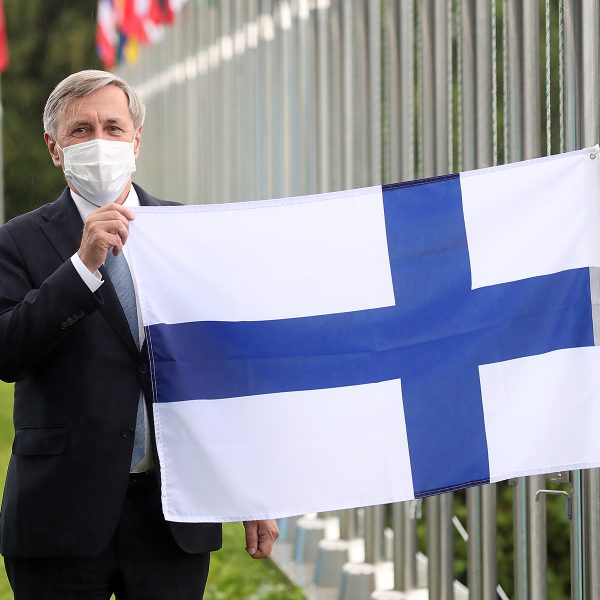
August 10, 2020 – SEOUL, South Korea – The flag of Finland was raised at the International Vaccine Institute (IVI) Headquarters today during a ceremony welcoming the country’s accession to IVI. Finland joined the Seoul-based international organization dedicated to vaccines for global health in […]
IVI hosted the “Shared Future, Global Solidarity: Vaccines Save Lives” event at its headquarters on July 8, 2020, attended by the First Lady, Minister of Health, Vice Minister of Foreign Affairs, Director of National Institute of Health of Korea, and foreign ambassadors to South Korea […]
LINE FRIENDS’ BT21 characters featured in IVI sticker set to raise funds for child immunization initiatives and COVID-19 vaccine development SEONGNAM, South Korea – June 30, 2020 – LINE Corporation today released a set of animated stickers together with the International Vaccine Institute (IVI) […]
June 16, 2020, SEOUL, South Korea – The International Vaccine Institute (IVI) welcomed His Excellency Einar H. Jensen, Ambassador of Denmark to South Korea, to IVI’s headquarters today to discuss developments in COVID-19 research as well as the institute’s ongoing studies on antimicrobial resistance […]
First COVID-19 vaccine clinical study approved in South Korea funded by CEPI through INOVIO, and supported by KCDC/KNIH 2-stage trial to test INOVIO’s COVID-19 vaccine (INO-4800) using well-established DNA platform technology in adults June 4, 2020, SEOUL, Korea and PLYMOUTH MEETING, PA, USA […]
June 3 2020, Oslo, Norway; Seoul, South Korea; Telangana, India—CEPI, the Coalition for Epidemic Preparedness Innovations, in collaboration with Ind-CEPI, has announced a new partnering agreement with a consortium comprising Bharat Biotech (BBIL) and the International Vaccine Institute (IVI) to advance the development of […]
SEOUL, KOREA — March 10, 2020 — Veeva Systems (NYSE:VEEV) today announced that the International Vaccine Institute (IVI), a not-for-profit International Organization established in 1997 as an initiative by the United Nations Development Programme (UNDP), has implemented Veeva Vault QualityDocs to improve control and real […]
May 22, 2020, SEOUL, South Korea – The International Vaccine Institute (IVI) invited French Ambassador to Korea, Philippe Lefort, to IVI’s headquarters in Seoul, Korea today to share ongoing research and development on COVID-19. His Excellency Philippe Lefort, French Ambassador to Korea (left), and […]
IVI will leverage its network of infectious disease surveillance sites to conduct epidemiological studies of COVID-19 in Madagascar and Burkina Faso Sida’s contribution will significantly build in-country capacity to proactively respond to the pandemic May 21, 2020 – SEOUL, South Korea – The […]
INO-4700 (GLS-5300) DNA vaccine demonstrates 100% binding and 92% neutralizing antibody responses against MERS-CoV INO-4800 DNA vaccine for COVID-19 currently in Phase 1 trial utilizes identical strategy targeting Spike protein and CELLECTRA intradermal delivery PLYMOUTH MEETING, Pa. and SEOUL, KOREA – April 28, 2020 […]
LINE to offer an Official Account to IVI to promote the importance of vaccines and vaccination, as well as feature educational contents starring IVI Goodwill Ambassador Henry Lau LINE FRIENDS’ BT21 character IVI sticker set to be launched in the first half of the year […]
The Coalition for Epidemic Preparedness Innovations (CEPI) grants funding $6.9 million to INOVIO and IVI to conduct clinical testing in Korea for INOVIO’s COVID-19 vaccine candidate based on their well-established DNA platform technology Korea National Institute of Health (KNIH) to support IVI’s testing efforts […]
The Asia Pacific region is vulnerable to the emergence and spread of AMR but there is little high-quality data available on the extent of its impact Quality-assured data is essential for building tailored strategies for preventing the spread of drug-resistant infections IVI is part of […]
January 13, 2020 – SEOUL, South Korea – The International Vaccine Institute (IVI), a Seoul, Korea-based international organization, announced today that the Swedish International Development Cooperation Agency (Sida) will continue to support IVI’s mission to accelerate vaccine research and development for global health […]
January 9, 2020 – SEOUL, South Korea – The International Vaccine Institute (IVI) received a $1.4 million grant from the Bill & Melinda Gates Foundation to ensure critical standards and reagents are available to low-cost oral cholera vaccine (OCV) manufacturers in the global health […]
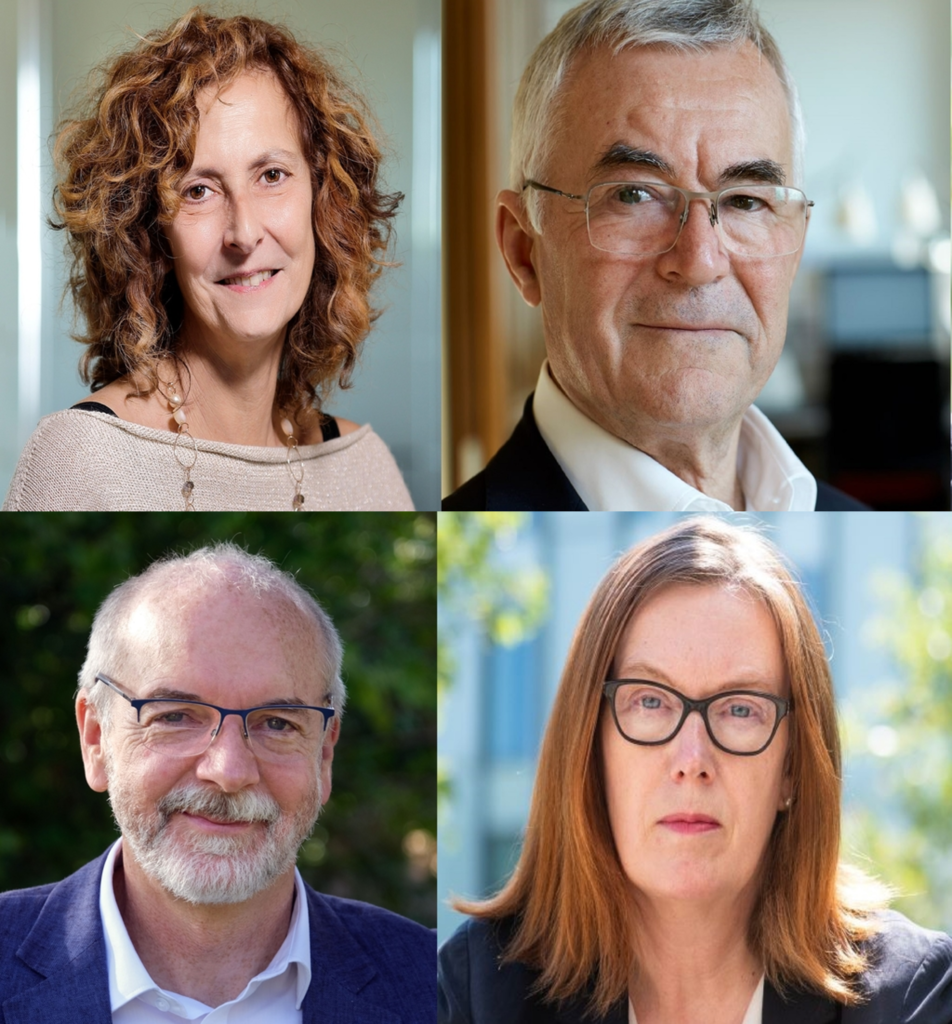
December 18, 2019 – SEOUL, South Korea – The International Vaccine Institute (IVI), a Seoul, Korea-based international organization dedicated to accelerating vaccines for global health, announced today that Professor Gordon Dougan, Dr. Melanie Saville, and Dr. Jean-Marie Okwo-Bele will join its Board of Trustees (BOT) […]
An award from the Wellcome Trust’s Affordable Innovations for Global Health Flagship will support IVI’s development and selection of potential vaccine candidates There is currently no vaccine available to protect against invasive non-typhoidal salmonella (iNTS) November 13, 2019 – SEOUL, South Korea – The […]
Institute expected to increase contributions to vaccine development activities worldwide November 1, 2019 – SEOUL, South Korea – The International Vaccine Institute (IVI), an international organization for vaccine development and delivery headquartered in Seoul, has been granted Good Clinical Laboratory Practice (GCLP) certification by […]
October 28, 2019 – SEOUL, South Korea – The International Vaccine Institute (IVI) has received a $4.5 million grant from the Bill & Melinda Gates Foundation to simplify the current oral cholera vaccine (OCV) and further increase its accessibility. To date, the Gates Foundation has […]
IVI State Forum 2019 was attended by 8th UN Secretary-General Ban Ki-moon, as well as 30 distinguished guests, including 17 ambassadors and top diplomats in Seoul, and senior Korean government officials. The IVI State Forum 2019 was attended by ambassadors and senior diplomats from […]
September 24, 2019, SEOUL, Korea – The Chief Medical Officer for England, Dame Sally Davies, announced today during the 74th UN General Assembly that a consortium led by the International Vaccine Institute (IVI) has been awarded a £2.7m Fleming Fund Regional Grant to improve data […]
19th IVI International Vaccinology Course brings together 175 trainees and faculty members from 49 countries The annual course has trained over 1,544 healthcare professionals in vaccine development, evaluation, production and policy to help increase developing nations’ capacity in vaccine research and immunization. September […]
August 28, 2019 – SEOUL, South Korea – The International Vaccine Institute (IVI) has received additional funds of $4.33 million from the Bill & Melinda Gates Foundation to expand Severe Typhoid in Africa (SETA) Program activities in close alignment with the “Effect of a […]
August 16, 2019 – SEOUL, South Korea – Dr. Jerome Kim, Director General of the International Vaccine Institute (IVI), joins the Human Vaccines Project as one of four distinguished global leaders to help push forward its effort to solve one the greatest public health challenges […]
Fund to support IVI’s projects to vaccinate children in developing countries throughout Asia-Pacific and Africa, to develop vaccines against emerging viruses Prof. Park Sang-chul, President of the Korea Support Committee for IVI, Prof. Sung Young-chul with his donation pledge and IVI Director General Dr. […]
Streptococcus pyogenes bacteria, which causes scarlet fever and other infections 30 May 2019, SEOUL, South Korea – The International Vaccine Institute (IVI) and Australia’s Murdoch Children’s Research Institute (MCRI) will coordinate a global push to free the world of Group A Streptococcus (Strep A), […]
SEOUL, South Korea — An international consortium, led by the International Vaccine Institute (IVI), has received funding from the UK’s Fleming Fund Regional Grants to conduct the Capturing Data on Antimicrobial Resistance (AMR) Patterns and Trends in Use in Regions of Asia (CAPTURA) project. The […]
– Grant from the Bill & Melinda Gates Foundation will support additional supply of novel TCV at affordable cost for public sector markets SEOUL, Korea –The International Vaccine Institute (IVI) announces receipt of a $15.7 million-dollar grant from the Bill & Melinda Gates Foundation […]
– Event organized by IVI to discuss ways of cooperation to help increase Korea’s vaccine self-sufficiency, and accelerate globalization of the Korean vaccine industry Key participants pose for a commemorative photo at the ‘IVI Forum for Enhancement of Cooperation’ at the National Assembly of the […]
Recently, the sensitivity of fecal microbiological cultures for detecting cholera has come under question. Researchers reporting in PLOS Neglected Tropical Diseases investigated this claim using a ‘vaccine probe’ analysis of a completed cholera vaccine cluster randomized trial to support the sensitivity of conventional microbiological culture […]
KPA, as group of experts in pediatric infectious diseases, to collaborate in academic research and projects on infectious disease prevention To support vaccination of children in developing countries through the ‘One for Three’ campaign, in partnership with IVI The International Vaccine Institute (IVI), […]
SEOUL, Korea — The International Vaccine Institute (IVI) has been awarded a $3,238,974 grant from the Bill & Melinda Gates Foundation to provide technical assistance support for studies to measure the effectiveness of typhoid conjugate vaccine (TCV) in West Africa. This grant comes after […]
– Bilateral ‘Master Implementing Partner Services Agreement’ expected to enable IVI to provide technical capabilities to CEPI Oslo, Norway; Seoul, Republic of Korea 11 February 2019—The Coalition for Epidemic Preparedness Innovations (CEPI) and the Republic of Korea-based International Vaccine Institute (IVI), an international organisation […]
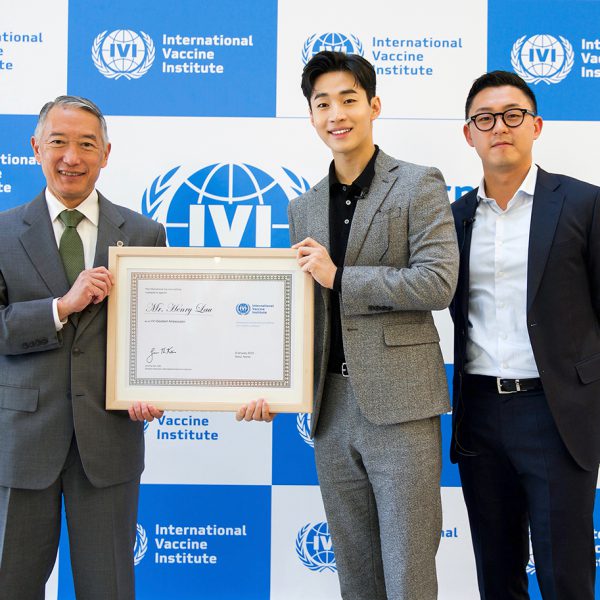
– Singer-songwriter, Henry to team up with the International Vaccine Institute (IVI), founded by the United Nations Development Programme to develop and deliver new vaccines for global health – IVI Director General Dr. Jerome Kim: “IVI is privileged to have Henry on board, and grateful […]
New study in Nature Communications reveals high burden of multi-drug resistant (MDR) typhoid fever in sub-Saharan Africa associated with two predominant genotypes – IVI epidemiologist Se Eun Park and collaborating scientists report data from Typhoid Fever Surveillance in Africa (TSAP) program and other sources […]
First cooperative program between International Vaccine Institute and Translational Health Science and Technology Institute expected to help accelerate cooperation between India and IVI IVI and the Translational Health Science and Technology Institute (THSTI) of India held a joint symposium at IVI headquarters on November 22. […]
“The Euvichol Story – Development and licensure of a safe, effective and affordable oral cholera vaccine through global public private partnerships” spearheaded by the International Vaccine Institute (IVI), has been published in Vaccine by Elsevier. The Euvichol Story published on October 9, 2018 in the […]
– Institute jointly conducts phase 1/2a clinical trial of GeneOne Life Science’s GLS-5300 DNA MERS vaccine – IVI in search of additional promising candidate vaccines among developers worldwide, with support from Samsung Life Public Welfare Foundation The International Vaccine Institute (IVI) said on September […]
Annual Course to update on the recent development in vaccinology, including basics of immunology, epidemiology, vaccine design and development Featured experts include: Stanley Plotkin (University of Pennsylvania), Dr. Barney Graham (US NIH), Dr. Randy Schoepp (USAMRID), Dr. Andy Pollard (University of Oxford), and Dr. Eric […]
KOICA, IVI team up with Mozambican MoH and INS, partners to vaccinate 190,000 people against cholera
– Joint initiative aims to prevent and control cholera, and secure sustainable cholera and diarrheal disease surveillance – WASH campaign to promote access to clean water and behavioral change for appropriate sanitation and hygiene practice – “KOICA is joining the program to contribute to ‘Ending […]
– Grant from the Bill & Melinda Gates Foundation to allow SK Bioscience Co. Ltd, IVI to accelerate late-stage development of new vaccine necessary to achieve WHO prequalification The International Vaccine Institute (IVI), an international nonprofit organization devoted to providing vaccines critical to global public […]
EuBiologics’ simplified OCV achieves WHO PQ
IVI starts technology transfer to Biological E. Limited to manufacture oral cholera vaccine for India and global markets
Prof. Jan Holmgren, and Profs. Barney Graham / Jason McLellan to receive the 3rd IVI-SK bioscience Park MahnHoon Award
Major typhoid fever surveillance study in sub-Saharan Africa indicates need for the introduction of typhoid conjugate vaccines in endemic countries
IVI participates as local coordinator in EID Conference & International Symposium for IDRIC
IVI opens a new Country and Project Office in Kenya that will lead the AVEC Africa project, working with partners and stakeholders on end-to-end vaccine R&D to ensure sustainable vaccine manufacturing in Africa
IVI to open Africa Regional Office in Rwanda
SK bioscience’s typhoid conjugate vaccine achieves WHO prequalification
Euvichol-S, simplified formulation of oral cholera vaccine, licensed by Korean regulatory agency
IVI and Gorgas Institute advocate for chikungunya vaccines at global meeting in Panama
CEPI, IVI and MRC Unit The Gambia partner to bolster clinical research capacity in West Africa to combat regional viral threats
CEPI-backed Research Preparedness Program West Africa aims to bolster regional clinical trial capacity and disease outbreak readiness. Partnership will support regional stakeholders to strengthen clinical research capacity to conduct Phase 2b / 3 clinical trials of Lassa fever vaccines in the region, for the region; […]
IVI, KAIST to collaborate in global vaccine research to accelerate innovations in vaccines and immunology
Joining forces in four main areas including vaccine immune response analysis using KAIST’s technology The International Vaccine Institute (IVI) and the Korea Advanced Institute of Science and Technology (KAIST) exchanged a memorandum of understanding (MOU) for global vaccine research collaboration at KAIST headquarters in […]
IVI signs MOU with University of Cambridge, University of Hong Kong, and the Hong Kong Jockey Club to establish the Hong Kong Jockey Club Global Health Institute
New typhoid conjugate vaccine Bio-TCV® approved in Indonesia
IVI conducts ‘2023 Introductory Course for Standard Practice (GxP Course)’ to support LMICs in local biomanufacturing
IVI, KEMRI-Wellcome Trust Research Programme, and Korea Biopharmaceutical CMO organize vaccine manufacturing training in Korea
IVI held an opening ceremony at headquarters in Seoul on October 30, 2023, to kickstart Hands-on Training for Upstream Process in Vaccine Manufacturing in partnership with KEMRI-Wellcome Trust Research Programme (KWTRP) and Korea Biopharmaceutical CMO (K-Bio CMO). Credit: IVI October 30, 2023 – SEOUL, […]
IVI conducts ‘KOR-IDB Biomanufacturing Training’ jointly with K-Bio Health and Korea’s leading manufacturers
IVI convenes inaugural Global Council meeting in Seoul
IVI’s 22nd International Vaccinology Course highlights lessons for vaccinology in the wake of the COVID-19 pandemic
IVI and Grid Biosciences partner for Epstein-Barr virus vaccine development with a commitment to global access
September 14, 2023, HONG KONG SAR and SEOUL, Republic of Korea – The International Vaccine Institute (IVI), an international organization with a mission to discover, develop, and deliver safe, effective, and affordable vaccines for global health, and Grid Biosciences (Grid), a biotech company developing […]
IVI and EuBiologics sign memorandum of understanding with DEK Vaccines Limited to support fill and finish of oral cholera vaccine in Ghana
IVI and CEPI renew partnership to accelerate development of vaccines against emerging infectious disease threats
IVI and the Madagascar Institute for Vaccine Research launch typhoid conjugate vaccine campaign for infants and children
Fiji launches mass vaccination campaign against typhoid in the Northern Division
SK bioscience’s COVID-19 vaccine makes WHO’s Emergency Use Listing
IVI exchanges MOU with Seoul National University on cooperation in research and beyond
June 21, 2023, SEOUL, Republic of Korea – The International Vaccine Institute (IVI) and Seoul National University signed an agreement to expand collaboration in research. IVI is the only international organization dedicated to the development and delivery of new vaccines to protect people […]
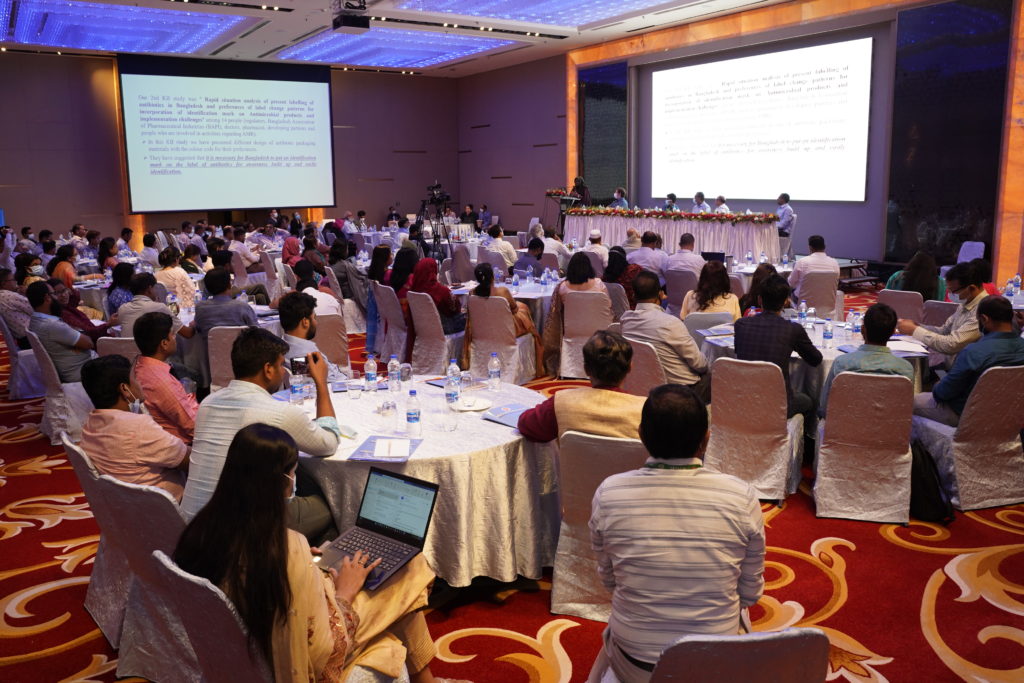
IVI-led CAPTURA consortium releases first country report on AMR data in Bangladesh
Report contains key findings for stakeholders engaging in AMR surveillance, research, policy, regulatory decision-making, and other infectious disease prevention and control programs in Bangladesh and wider Asia, supported by the Fleming Fund. June 20, 2023, SEOUL, Republic of Korea – CAPTURA is a […]
IVI conducts 2nd Introductory Course for Biologics Development and Manufacturing for 235 trainees from 47 LMICs
2-week training focuses on technology transfer, talent development, facility management, and aseptic processing Part of the Korean MOHW’s Global Training Hub for Biomanufacturing, the program aims to strengthen local bio production capabilities in LMICs to address vaccine inequity and strengthen pandemic preparedness June […]
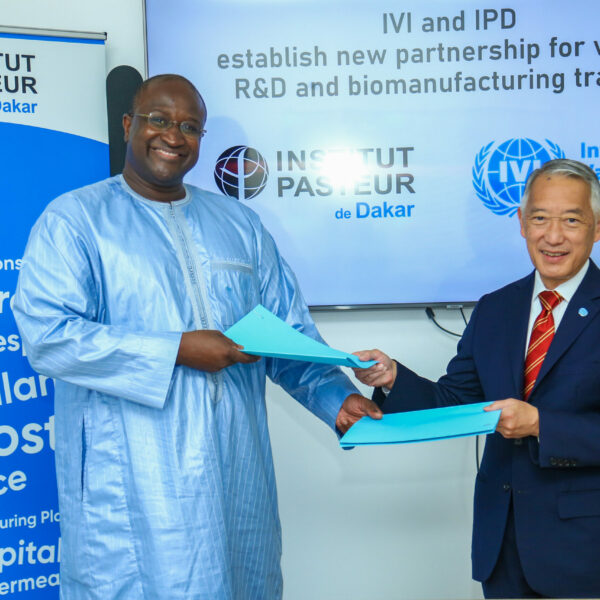
IVI and Institut Pasteur de Dakar establish new partnership for vaccine R&D and biomanufacturing training
SK bioscience’s COVID-19 Vaccine receives Marketing Authorization from UK regulatory authority
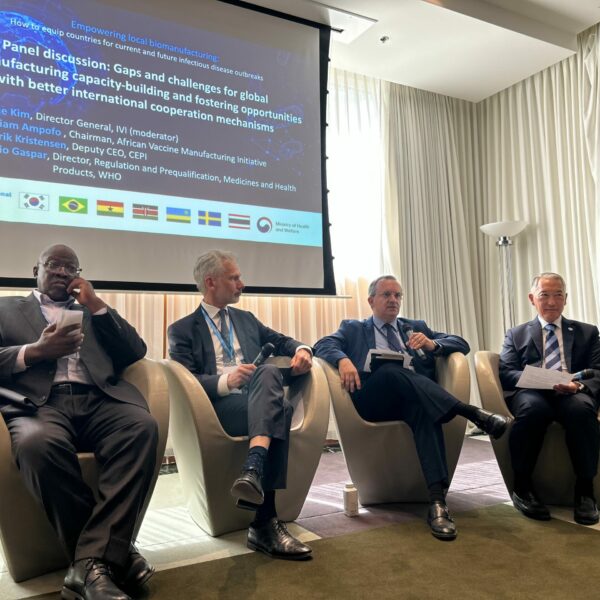
IVI and the Ministry of Health and Welfare of the Republic of Korea urge global action on enhancing vaccine manufacturing during the 76th World Health Assembly
IVI begins clinical development of DuoChol oral cholera vaccine
Drs. Rino Rappuoli and Mariagrazia Pizza; Profs. Andrew Pollard and Sarah Gilbert honored at 2023 IVI-SK bioscience Park MahnHoon Award Ceremony
Awardees give lectures to highlight their work and achievements, share knowledge at the Award Forum April 26, 2023 – SEOUL, Korea – The International Vaccine Institute (IVI) gave the 2023 IVI-SK bioscience Park MahnHoon Award to Drs. Rino Rappuoli and Mariagrazia Pizza as […]
IVI, Seegene exchange MOU on global HPV burden study
Seegene to handle logistics support to test 50,000 girls and women ages 9-50 in Asia and Africa from August Seegene’s Allplex™ HPV28 Detection tests to be used to improve access to HPV screening Project funded by the Bill & Melinda Gates Foundation, conducted jointly by […]
IVI welcomes Panama as a State Party
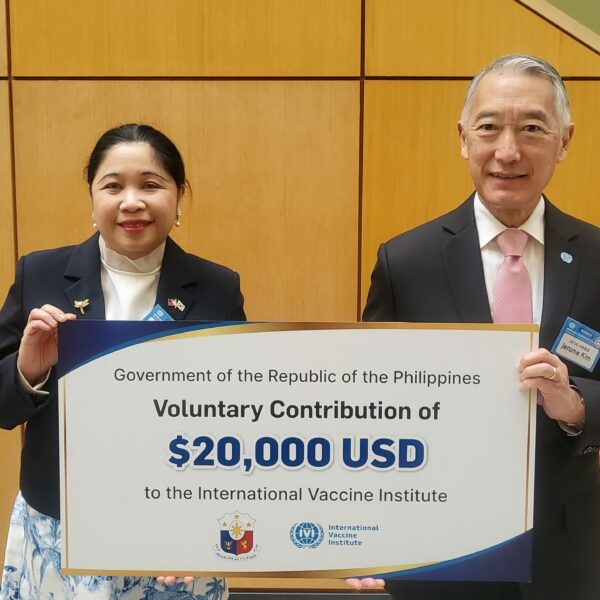
Government of the Republic of the Philippines makes voluntary contribution to IVI
March 14, 2023 – SEOUL, Republic of Korea – The Philippine Ambassador to the Republic of Korea, H.E. Ma. Theresa Dizon-de Vega, announced that the Government of the Republic of the Philippines has made a voluntary contribution to the International Vaccine Institute, an international […]
First Lady Kim Keon Hee inaugurated as 5th Honorary President of IVI Support Committee
Profs. Andrew Pollard / Sarah Gilbert and Drs. Rino Rappuoli / Mariagrazia Pizza named winners of second IVI – SK bioscience Park MahnHoon Award
IVI launches global study to determine the burden of HPV among girls and women
IVI opens Country Office in Austria
Gachon University, Korea mRNA Vaccine Initiative, IVI exchange MOU for joint vaccine R&D and mutual cooperation
IVI, Batavia Biosciences exchange memorandum of understanding for vaccine research and development
IVI conducts on-site training, consultation for Afrigen in South Africa
Provides training for about 40 Afrigen personnel on overall biopharmaceutical production as part of GTH-B program December 13, 2022, SEOUL, Republic of Korea — The International Vaccine Institute (IVI) conducted on-site training and consultation for Afrigen Biologics & Vaccines (Afrigen) in South Africa to help […]
IVI honored with Industry Minister’s Prize for Biosecurity Management
Clover’s Vaccine Candidate Reduced Household Transmission of SARS-CoV-2 in Study Published in Clinical Infectious Diseases
IVI, ST Pharm exchange MOU on clinical development of COVID-19 vaccine
Biovac signs deal with IVI to develop and manufacture oral cholera vaccine for African and global markets
IVI partners with Global Vaccine Leading Technology Center to accelerate vaccine development for global health
SK bioscience makes donation to IVI to support global R&D
Lemonex, IVI exchange MOU on joint vaccine development
November 10, 2022, SEOUL, Republic of Korea – The International Vaccine Institute (IVI) announced on November 10 it signed a memorandum of understanding (MOU) with Lemonex, a company specializing in RNA gene therapy development, and agreed to seek cooperation in research and development. IVI and Lemonex […]
IVI conducts 1st Introductory Course for Standard Practice (GxP Course) Oct. 31 – Nov. 18
IVI welcomes Thailand as a state party
IVI gathers member countries for 2022 State Forum and celebrates its 25th anniversary with key partners
IVI launches Global Advisory Group of Experts chaired by Dr. Deborah Birx
IVI signs memorandum of understanding with Hilleman Laboratories to advance vaccine R&D and manufacturing for safe, effective, and affordable vaccines in low-and middle-income countries
IVI and the London School of Hygiene & Tropical Medicine partner to accelerate vaccines for global health through joint training and research and development
IVI’s 21st International Vaccinology Course highlights breakthroughs in vaccine science and technology
IVI’s Europe Regional Office begins operations in Stockholm
September 1, 2022, SEOUL, Republic of Korea – The International Vaccine Institute (IVI), an international organization with a mission to discover, develop, and deliver safe, effective, and affordable vaccines for global health, opened the IVI Europe Regional Office (IERO) in Stockholm today. IERO, housed temporarily […]
Danish Minister for Health visits the International Vaccine Institute during official visit to Republic of Korea
First ‘Introductory Course for Biologics Development and Manufacturing’ completes training of 106 participants from 24 countries, and 32 Koreans
IVI and Moderna sign memorandum of understanding for vaccine research and development
IVI kicks off first ‘Introductory Course for Biologics Development and Manufacturing’ to train 117 people from 25 LMICs on July 18
Spain joins IVI with flag-raising ceremony
Typhoid: Multi- and single-dose formulations of Vi-DT vaccine shown to be safe and immunologically equivalent in new study
New evidence supports the use of alternative dosing formulations, expanding TCV delivery options in public health programs June 14, 2022 – SEOUL, Republic of Korea – A new study jointly conducted by the International Vaccine Institute (IVI) and collaborators shows multidose and single-dose […]
International Vaccine Institute and Egyptian Agency of Partnership for Development co-host symposium on African vaccine manufacturing
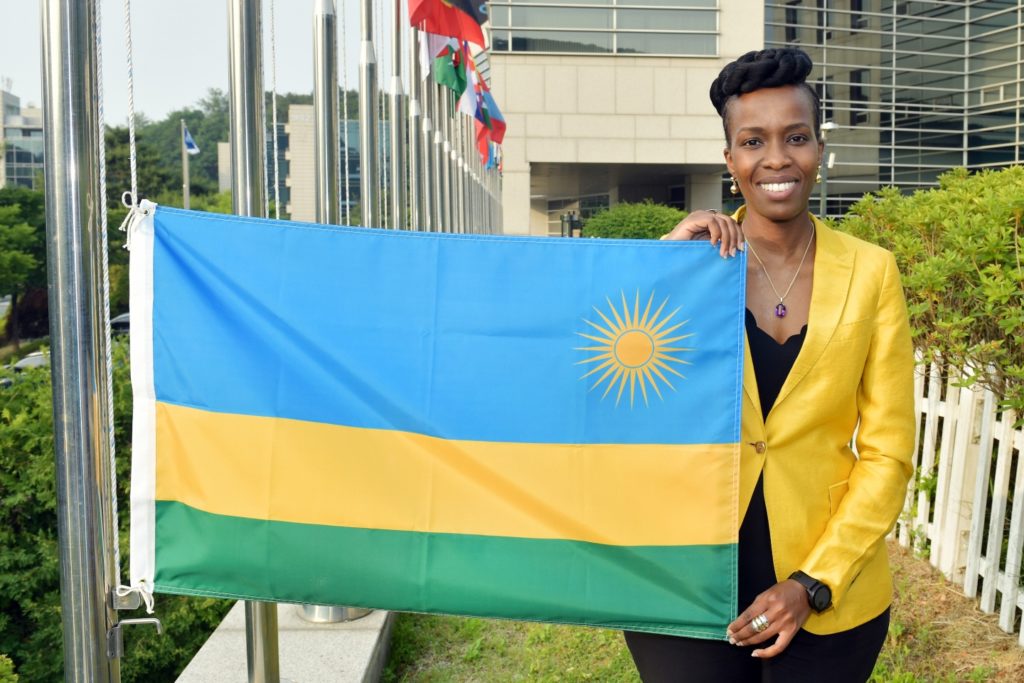
IVI welcomes Rwanda as a member state with flag-raising ceremony
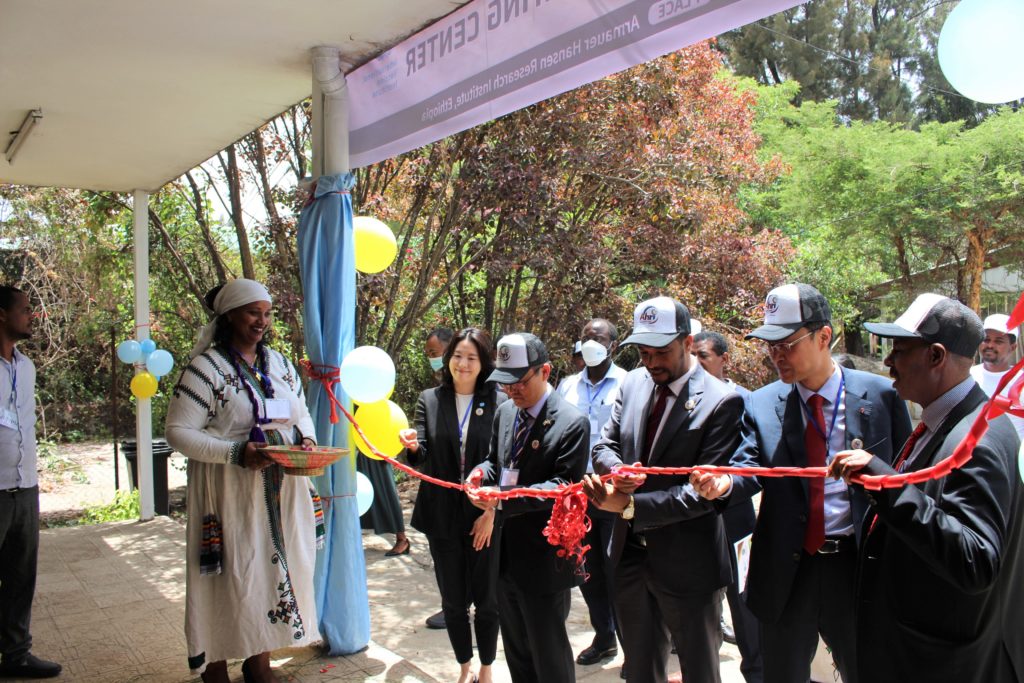
IVI and AHRI establish Collaborating Center for joint vaccine research, development, and capacity-building for global health
May 23, 2022– SEOUL, Republic of Korea and ADDIS ABABA, Ethiopia – The International Vaccine Institute (IVI), an international organization with a mission to discover, develop, and deliver safe, effective, and affordable vaccines, and Armauer Hansen Research Institute (AHRI), established by the Government of […]
IVI and the Government of Bangladesh join forces to boost AMR response in South Asia
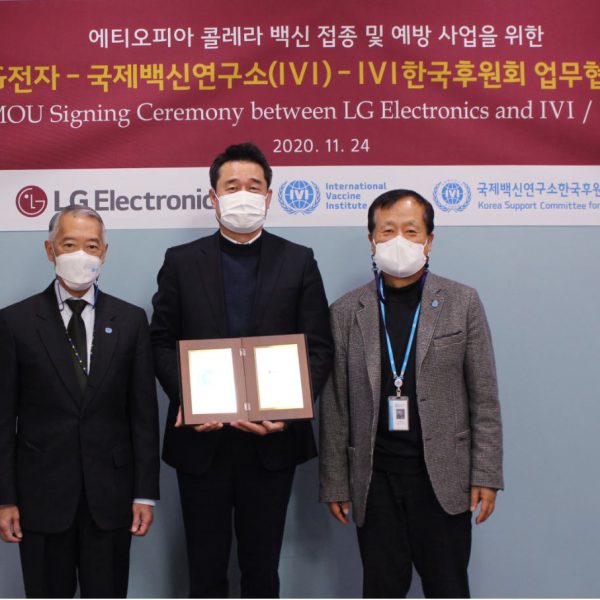
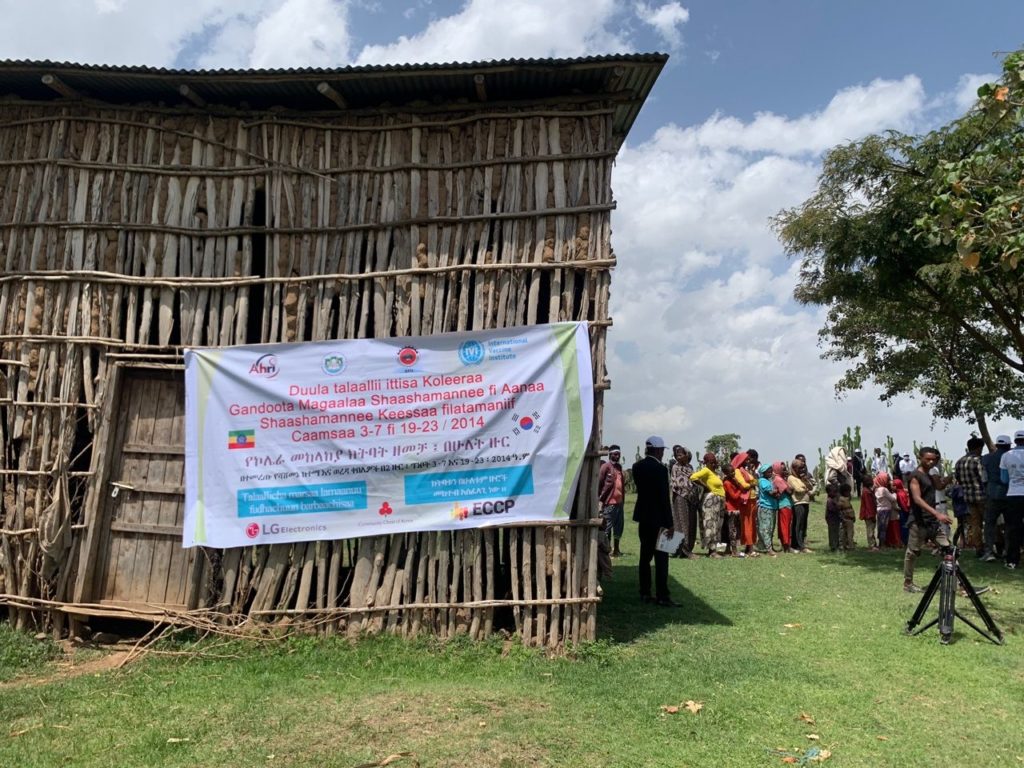
IVI, LG Electronics, Korean donors, AHRI and EPHI join hands to vaccinate 100,000 people at risk of cholera in Ethiopia
IVI, KDCA exchange MOU on ROK’s state funding, strengthened cooperation
New typhoid conjugate vaccine licensed in Korea
IVI and SK bioscience confirm SK’s COVID-19 vaccine candidate meets coprimary objectives in a joint 6-country Phase III study
Recombinant protein-based ‘SKYCovione™’ adjuvanted with GSK’s pandemic adjuvant demonstrates superior neutralizing titers compared to a control vaccine and safety SK bioscience submits a biologics license application to KMFDS, and will submit license applications to international regulatory agencies Company set to supply 10 million doses of […]
IVI, SmileGate, partners team up to vaccinate 28,000 people in Nepal
2022 IVI-SK bioscience Park MahnHoon Award Ceremony honors Dr. Tore Godal, Profs. Drew Weissman & Katalin Karikó
International Vaccine Institute appoints two members to its Board of Trustees representing Ecuador and Rwanda
IVI, ROK’s MOHW expand partnership to promote vaccine development & delivery, bio manufacturing training for LMICs
Dr. Tore Godal, Profs. Drew Weissman & Katalin Karikó honored with first IVI – SK bioscience Park MahnHoon Award
IVI welcomes the United Arab Emirates as its 37th member state with flag-raising ceremony
March 17, 2022 – SEOUL, South Korea – The International Vaccine Institute (IVI) announced today that the United Arab Emirates (UAE) joined IVI, becoming the international organization’s 37th member state. His Excellency Abdulla Saif Al Nuaimi, Ambassador of the UAE to the Republic of […]
IVI named operator of ‘2022 Global Bio-Intensive Training Courses’ by Korean Ministry of Health and Welfare
Institute to run training program on vaccines and biologics development and manufacturing and GxP for 450 trainees from LMICs and Korea February 16, 2022 – SEOUL, Republic of Korea – The International Vaccine Institute (IVI) has been designated by the Ministry of Health and Welfare […]
New studies: Vi-DT vaccine is safe and immunogenic, with booster dosing potentially eliciting long-term immune responses
January 27, 2022 – SEOUL, Republic of Korea – A new study shows that late booster dosing with Vi polysaccharide conjugated with diphtheria toxoid (Vi-DT), one of the typhoid conjugate vaccines (TCVs), at 27.5 months post-first dose is safe and elicits robust immune responses in […]
IVI and NVI partner for collaborative vaccine R&D and capacity building
IVI and the National Vaccine Institute (NVI) of Thailand signed a Definitive Agreement to strengthen their collaborative partnership January 20, 2022 – SEOUL, Republic of Korea – The International Vaccine Institute (IVI) and the National Vaccine Institute (NVI) of Thailand signed a Definitive […]
IVI and SK bioscience Complete Recruitment for Phase III Clinical Trial of SKBS’ COVID-19 Vaccine
Recruitment of global Phase III clinical trial of ‘GBP510’ completed, with vaccine expected to be authorized in the first half of 2022 A booster dose study is concurrently being planned to bring additional vaccine to the global COVID-19 vaccine market January 19, 2022 SEOUL, […]
IVI and Mozambique’s INS launch mix-and-match COVID-19 vaccine study
Study will assess the safety and immunogenicity of a heterologous regimen of two approved COVID-19 vaccines in adults Aims to mitigate vaccine shortages and support flexible immunization programs December 20, 2021, SEOUL, Republic of Korea; MAPUTO, Mozambique; OSLO, Norway – The International Vaccine Institute […]
IVI to establish a European regional office in Sweden
IVI and the Government Offices of Sweden signed a Memorandum of Understanding today to establish the office in Stockholm, creating a European hub for global health research and innovation December 17, 2021, SEOUL, Republic of Korea – The International Vaccine Institute (IVI), an international […]
IVI, ADB SEADS, IPK, ICARS, and the Danish Embassy in Korea launch webinar on combatting antimicrobial resistance
The webinar “Curbing the Invisible Pandemic: Effective Solutions to Collectively Combat Antimicrobial Resistance” was held on December 7, 2021 December 7, 2021 — The International Vaccine Institute (IVI), Asian Development Bank Southeast Asia Development Solutions (ADB SEADS), Institut Pasteur Korea (IPK), the […]
Dr. Sushant Sahastrabuddhe of IVI named ‘Honorary Citizen of Seoul’
Cited for contributions to development of vaccines, joining 8 foreign residents to receive Honorary Citizenship in 2021 Second Honorary Seoul Citizen from IVI SEOUL, Korea — Dr. Sushant Sahastrabuddhe, Associate Director General at the International Vaccine Institute (IVI), has been awarded “Honorary Citizenship of […]
IVI partners with SK bioscience to launch new prestigious award for vaccine industry
IVI, SK sign an agreement to launch the ‘Park MahnHoon Award’ to honor individuals and organizations that made contributions to the vaccine industry Award Ceremony will be held annually in April; IVI and SK ‘committed to establish the new award as a prestigious award for […]
IVI Director General named Honorary Ambassador of Korean Tourism
To promote the safety and attractiveness of traveling Korea amidst gradual restart of global tourism The Ministry of Culture, Sports and Tourism (Minister Hwang Hee) and the Korea Tourism Organization (KTO, President Ahn Young-bae) appointed Dr. Jerome H. Kim, Director General of the […]
IVI receives Development Cooperation Award from Republic of Korea
The International Vaccine Institute (IVI) has been honored with the Presidential Citation of the Republic of Korea’s Development Cooperation Award, in recognition of IVI’s contributions to global health and Korea’s official development assistance. IVI has been cited as being an “International organization focused […]
IVI leaders address health security at ‘first World Emerging Security Forum’ in Seoul
Dr. Jerome Kim, Director General and Dr. Song Manki, Deputy Director General of Science of the International Vaccine Institute (IVI) joined distinguished global leaders and experts as moderator and speaker at the first World Emerging Security Forum, which took place in Seoul on November […]
IVI and JEDI partner for innovations in health and science
October 25, 2021 – SEOUL, Republic of Korea, PARIS, France, BRUSSELS, Belgium, BERLIN, Germany – The International Vaccine Institute (IVI) and the Joint European Disruptive Initiative (JEDI) signed a Memorandum of Understanding to establish a collaborative relationship. Both organizations are dedicated to advancing innovations in […]
IVI’s 2021 State Forum highlights opportunities to bridge global vaccine gaps
Until Everyone Is Safe: Global Vaccine Needs and IVI’s Capabilities was livestreamed on October 7, 2021 at 16:00 Korea Standard Time, and a recording of the event is available here. October 7, 2021, SEOUL, Republic of Korea — The International Vaccine […]
The International Vaccine Institute, Texas Children’s Hospital, and Baylor College of Medicine partner for collaborative vaccine research
Sept. 28, 2021 – SEOUL, South Korea, and HOUSTON, Texas, USA – The International Vaccine Institute (IVI), the Texas Children’s Hospital Center for Vaccine Development, and Baylor College of Medicine’s National School of Tropical Medicine signed a Memorandum of Understanding to promote and further academic […]
Enrollment begins for Sanofi and GSK’s Phase 3 efficacy trial of COVID-19 vaccine candidate in Nepal, led by IVI
International Vaccine Institute (IVI) will lead the clinical trial in Nepal to assess the safety, efficacy and immunogenicity of an adjuvanted recombinant-protein COVID-19 vaccine candidate Phase 3 international clinical trial includes volunteers from several countries, including sites in the US, Asia, Africa and Latin America […]
IVI’s 20th International Vaccinology Course brings virtual training to vaccine professionals and students across 155 countries
September 10, 2021, SEOUL, Korea — The International Vaccine Institute’s (IVI) 20th International Vaccinology Course (IVC) concluded today, bringing together 7,350 registered trainees and 27 faculty members for 5 days of online lectures on a range of topics related to the science of vaccines with […]
IVI 20th International Vaccinology Course kicks off September 6, 2021
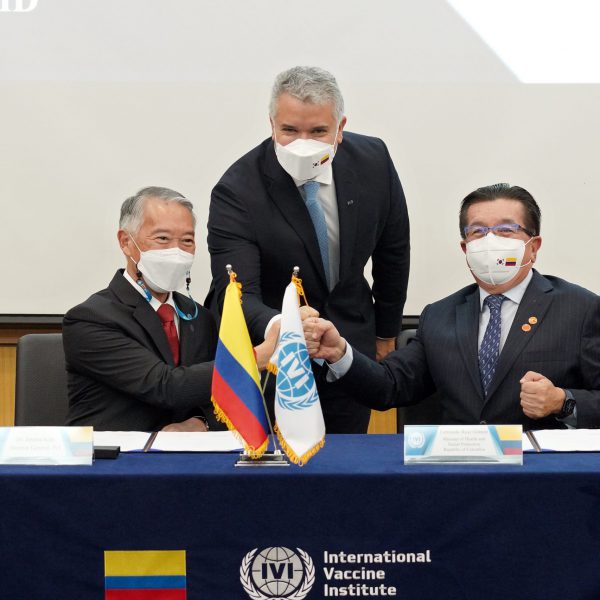
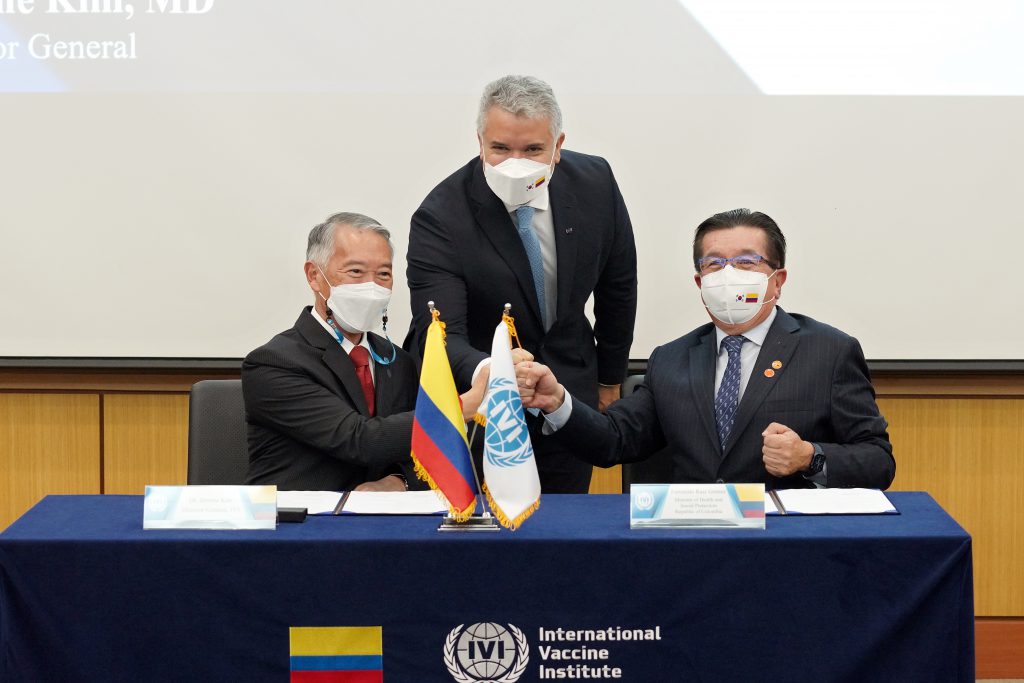
President Duque of Colombia visits the International Vaccine Institute
IVI and BBIL launch global Chikungunya vaccine Phase II/III trial in Costa Rica
IVI is leading the Global Chikungunya vaccine Clinical Development Program (GCCDP) consortium in partnership with Bharat Biotech International Ltd. and with support from the Coalition for Epidemic Preparedness Innovations (CEPI) and Ind-CEPI. August 24, 2021 – SEOUL, Republic of Korea – The International Vaccine […]
IVI appoints ballerina Sae Eun Park to IVI Goodwill Ambassador
August 18, 2021 – SEOUL, Republic of Korea – The International Vaccine Institute (IVI) named Sae Eun Park, an award-winning ballet dancer from Korea, an IVI Goodwill Ambassador during a virtual ceremony in Seoul today. Ms. Park was named danseuse étoile, the highest-level dancer, […]
IVI partners with SK bioscience to conduct late-stage global clinical trials of SK bioscience’s COVID-19 vaccine
THECA consortium begins Vi-TT typhoid conjugate vaccine phase IV effectiveness study in Ghana
July 26, 2021 – SEOUL, Republic of Korea – The International Vaccine Institute (IVI) announced today the start of a cluster-randomized, controlled Phase IV trial to assess the effectiveness of Typbar® typhoid conjugate vaccine (TCV) in preventing typhoid infection in children in Asante Akim, Ghana […]
CEPI and IVI collaborate on clinical research to expand access to COVID-19 vaccines in Africa
July 20, 2021; Oslo, Norway and Seoul, Republic of Korea: The Coalition for Epidemic Preparedness Innovations (CEPI) and the International Vaccine Institute (IVI) today announced a new programme of clinical research which aims to expand access to COVID-19 vaccines in Africa. CEPI will provide funding […]
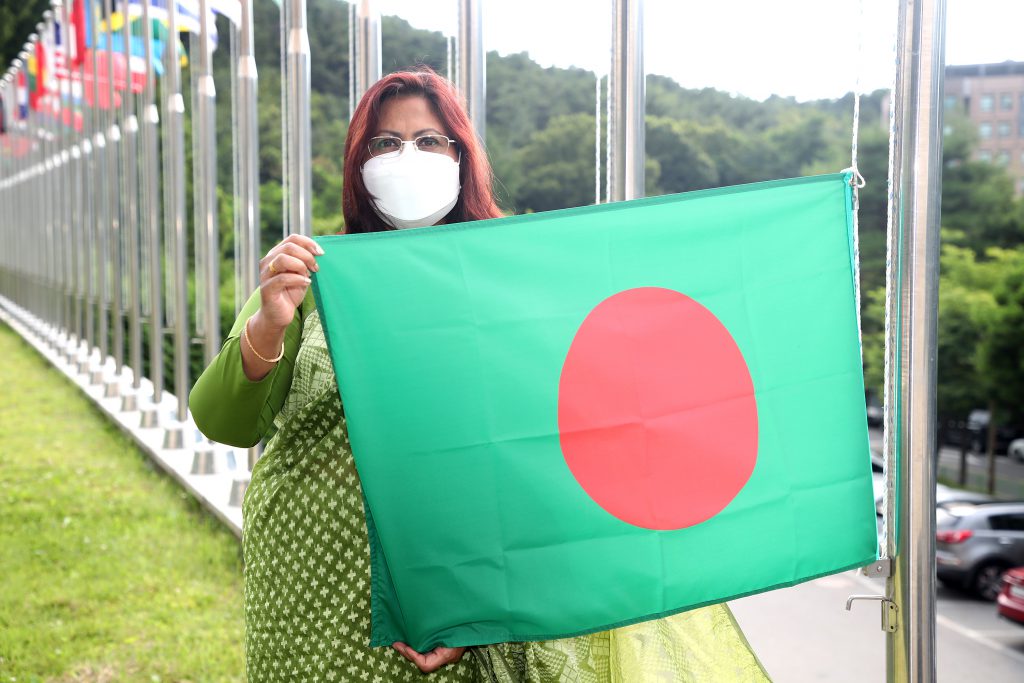
IVI welcomes Bangladesh as state party with ratification ceremony
July 15, 2021 – SEOUL, South Korea – The International Vaccine Institute (IVI) hosted a ceremony at headquarters today honoring the People’s Republic of Bangladesh’s ratification of the IVI Establishment Agreement. Over 20 years of vaccine research and capacity-building initiatives with partners in Bangladesh […]
IVI and KNUST establish collaborating center to conduct vaccine research and development for global health
IVI organizes ROK-Australia-ASEAN Vaccine Forum to promote inclusive and sustainable recovery from the COVID-19 pandemic
The ROK-Australia-ASEAN Vaccine Forum took place in hybrid live/virtual format with transmission from Seoul on June 29, 2021 June 29, 2021 – SEOUL, Republic of Korea – The International Vaccine Institute (IVI) organized, and the Republic of Korea’s Ministry of Foreign Affairs (MOFA) and […]
IVI to develop an adaptive Phase 1b/2a schistosomiasis vaccine clinical trial
May 31, 2021 – SEOUL, South Korea – The International Vaccine Institute (IVI) announced today that the Bill & Melinda Gates Foundation awarded a grant to IVI to develop an adaptive trial design protocol for a Phase 1b/2a clinical trial of a schistosomiasis vaccine. […]
International Vaccine Institute and FII Institute sign MOU to cooperate on scientific research and publication
The agreement is an effort to further scientific research in infectious diseases and make knowledge more accessible through advocacy and joint publications. May 19, 2021 – SEOUL, South Korea, RIYADH, Saudi Arabia – The International Vaccine Institute (IVI) and the Future Investment Initiative Institute […]
International Vaccine Institute and Institut Pasteur Korea Sign MOU for Research and Development of Infectious Disease Therapeutics and Vaccines
April 1, 2021, GYEONGDO-DO and SEOUL, Korea – The International Vaccine Institute (IVI) and the Institut Pasteur Korea (IPK) signed a memorandum of understanding (MOU) for mutual collaboration in the research and development of therapeutics and vaccines for infectious diseases. The MOU signing ceremony, […]
IVI signs Memorandum of Understanding with the Nepal Health Research Council
IVI welcomes Bangladesh’s accession to IVI
March 9, 2021 – SEOUL, South Korea – The International Vaccine Institute (IVI) welcomes the Cabinet of Bangladesh’s decision to ratify the IVI Establishment Agreement on February 22, 2021, completing the full accession process to IVI and becoming its 19th State Party. Bangladesh is a founding signatory to IVI’s […]
George Bickerstaff re-elected Chairperson of the International Vaccine Institute’s Board of Trustees
February 22, 2021, SEOUL, Korea – The International Vaccine Institute (IVI) announced today that its Board of Trustees (BOT) re-elected Mr. George Bickerstaff as Chairperson of the BOT. His second three-year term will begin this May. Mr. Bickerstaff has served on the IVI Board […]
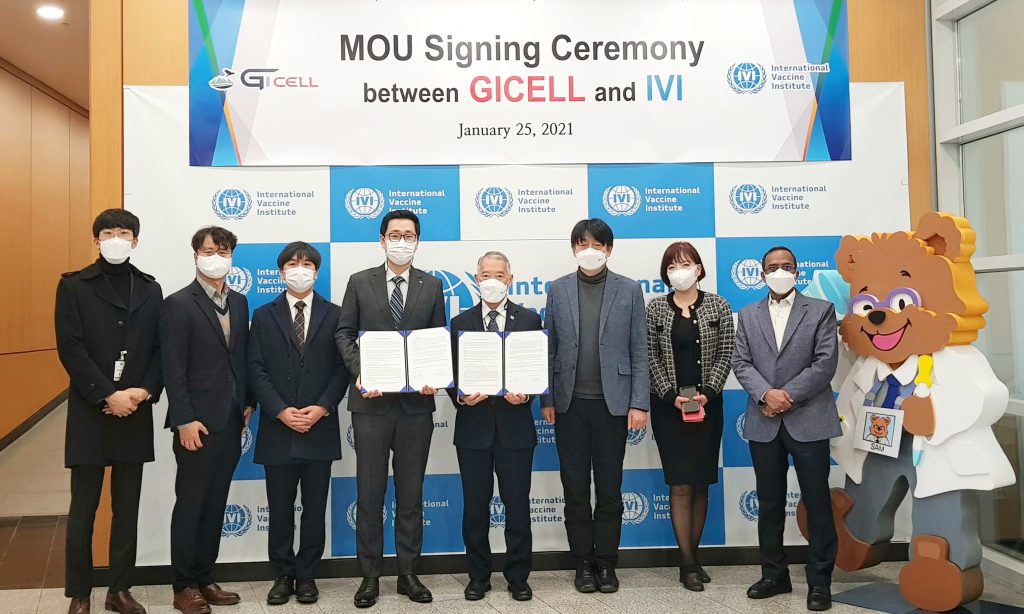
GI-Cell, IVI sign MOU to develop next-generation COVID-19 vaccine
Cellid, IVI exchange ‘COVID-19 Vaccine Research Service Contract’
IVI to analyze samples from Phase 1/2a trials of Cellid’s COVID-19 vaccine to evaluate immunogenicity January 5, 2021, SEOUL, Korea – Cellid and the International Vaccine Institute (IVI) have exchanged a collaborative research agreement to analyze the immunogenicity of the COVID-19 vaccine “AdCLD-CoV19”. […]
IVI-SK’s new typhoid conjugate vaccine meets primary endpoints in phase III study in Nepal
Primary analysis also confirms safety of Vi-DT December 17, 2020 – SEOUL, South Korea – Vi-DT typhoid conjugate vaccine, developed jointly by the International Vaccine Institute (IVI) and SK bioscience, has met the primary endpoints in a phase III study in Nepal. The primary […]
EuBiologics partners up with IVI in COVID-19 vaccine development
December 3, 2020, SEOUL, South Korea – The International Vaccine Institute (IVI) and EuBiologics exchanged an MOU to cooperate in clinical development of the COVID-19 vaccine the company is currently developing. The signing ceremony at IVI headquarters on December 2 was attended by Dr. Jerome […]
How to take action on AMR during the COVID-19 pandemic, a webinar co-hosted by IVI, ICARS, and the Embassy of Denmark in Korea
Free webinar, Evidence to Action: Advancing the Antimicrobial Resistance Agenda during a Pandemic, will be held on Thursday, December 3, 2020 at 9:00 Central European Time (17:00 Korean Standard Time). Registration available at ivi.int/evidence-to-action/ December 1, 2020, SEOUL, Korea — The International Vaccine Institute […]
IVI celebrates GivingTuesday with local community heroes and launches new donation initiative
December 1, 2020, SEOUL, South Korea – The International Vaccine Institute (IVI) visited the Seoul Gwanak Police Station today to recognize the Foreign Affairs section for their ongoing support for IVI’s activities and to deliver protective face masks and medical supplies for staff and volunteers. […]
IVI, ROK’s GDEF join forces to provide OCV vaccination to 540,000 people at risk of cholera in Nepal and Mozambique
GDEF’s US$8.05 million grant to support ECHO projects to prevent and control cholera and contribute to ‘Ending Cholera—A Global Roadmap to 2030’ The International Vaccine Institute (IVI) and the Republic of Korea’s Global Disease Eradication Fund (GDEF) have agreed to conduct joint projects to […]
IVI and the Ministry of Foreign Affairs and Human Mobility of Ecuador exchange MOU to pursue global health research & development
November 24, 2020, SEOUL, Republic of Korea — The International Vaccine Institute (IVI) and the Ministry of Foreign Affairs and Human Mobility of Ecuador exchanged a memorandum of understanding (MOU) today at IVI headquarters in Seoul, Republic of Korea to explore areas of collaboration in […]
LG Electronics, IVI join forces to conduct a ‘cholera vaccination and prevention project’ in Ethiopia
Project to provide vaccination for 40,000 residents in areas at risk of cholera and establish disease monitoring system with the Ethiopian Ministry of Health through the Armauer Hansen Research Institute (AHRI) To contribute to health authorities in policymaking for disease prevention by investigating waterborne diseases […]
IVI, Vaccine Innovative Technology ALliance Korea (VITAL-Korea) to partner up for innovative vaccine research and development
Aim to accelerate R&D and globalization of Korean vaccines to increase contributions to global health The International Vaccine Institute (IVI) and the Vaccine Innovative Technology ALliance Korea (VITAL-Korea) agreed to join forces to promote vaccine research and development for global health. IVI […]
International Vaccine Institute honored with Minister of Health and Welfare Award for achievements in biosafety
October 28, 2020, SEOUL, Korea — The International Vaccine Institute (IVI) received an award by the Minister of Health and Welfare of the Republic of Korea for achievements in biosafety management. IVI was honored with the Minister of Health and Welfare Award for achievements in […]
The International Vaccine Institute Supports a Global Campaign to Reduce the Spread of Covid-19
The International Vaccine Institute (IVI) is partnering with the World Sanity Foundation (WSF) in a campaign to encourage the use of face masks in an effort to reduce the spread of COVID-19 cases until a safe and effective vaccine becomes widely available. The first […]
IVI and global health partners encourage vaccine diplomacy
Typhoid: Study confirms Vi-DT conjugate vaccine is safe and immunogenic in children 6-23 months of age
IVI to ready clinical trial sites for COVID-19 vaccine efficacy trials in 4 countries
IVI welcomes Finland as 36th member state with flag-raising ceremony
August 10, 2020 – SEOUL, South Korea – The flag of Finland was raised at the International Vaccine Institute (IVI) Headquarters today during a ceremony welcoming the country’s accession to IVI. Finland joined the Seoul-based international organization dedicated to vaccines for global health in […]
IVI welcomes the First Lady of South Korea to a solidarity event for global health and vaccine equity
IVI hosted the “Shared Future, Global Solidarity: Vaccines Save Lives” event at its headquarters on July 8, 2020, attended by the First Lady, Minister of Health, Vice Minister of Foreign Affairs, Director of National Institute of Health of Korea, and foreign ambassadors to South Korea […]
LINE and International Vaccine Institute Release BT21 Donation Stickers to Promote Global Vaccination and Vaccine Development
LINE FRIENDS’ BT21 characters featured in IVI sticker set to raise funds for child immunization initiatives and COVID-19 vaccine development SEONGNAM, South Korea – June 30, 2020 – LINE Corporation today released a set of animated stickers together with the International Vaccine Institute (IVI) […]
IVI welcomes H.E. Einar H. Jensen, Ambassador of Denmark to South Korea, to discuss COVID-19 and AMR research
June 16, 2020, SEOUL, South Korea – The International Vaccine Institute (IVI) welcomed His Excellency Einar H. Jensen, Ambassador of Denmark to South Korea, to IVI’s headquarters today to discuss developments in COVID-19 research as well as the institute’s ongoing studies on antimicrobial resistance […]
IVI partners with Seoul National University Hospital to start Phase 1/2 clinical trial of INOVIO’s COVID-19 DNA (INO-4800) vaccine in South Korea
First COVID-19 vaccine clinical study approved in South Korea funded by CEPI through INOVIO, and supported by KCDC/KNIH 2-stage trial to test INOVIO’s COVID-19 vaccine (INO-4800) using well-established DNA platform technology in adults June 4, 2020, SEOUL, Korea and PLYMOUTH MEETING, PA, USA […]
CEPI awards up to US$14.1million to consortium of IVI and Bharat Biotech to advance development of Chikungunya vaccine in collaboration with Ind-CEPI
June 3 2020, Oslo, Norway; Seoul, South Korea; Telangana, India—CEPI, the Coalition for Epidemic Preparedness Innovations, in collaboration with Ind-CEPI, has announced a new partnering agreement with a consortium comprising Bharat Biotech (BBIL) and the International Vaccine Institute (IVI) to advance the development of […]
IVI standardizes on Veeva Vault QualityDocs to Improve Operational Efficiency
SEOUL, KOREA — March 10, 2020 — Veeva Systems (NYSE:VEEV) today announced that the International Vaccine Institute (IVI), a not-for-profit International Organization established in 1997 as an initiative by the United Nations Development Programme (UNDP), has implemented Veeva Vault QualityDocs to improve control and real […]
IVI presents COVID-19 vaccine and epidemiology research to H.E. Philippe Lefort, Ambassador of France to Korea
May 22, 2020, SEOUL, South Korea – The International Vaccine Institute (IVI) invited French Ambassador to Korea, Philippe Lefort, to IVI’s headquarters in Seoul, Korea today to share ongoing research and development on COVID-19. His Excellency Philippe Lefort, French Ambassador to Korea (left), and […]
IVI to strengthen COVID-19 surveillance in sub-Saharan Africa with support from Sweden
IVI will leverage its network of infectious disease surveillance sites to conduct epidemiological studies of COVID-19 in Madagascar and Burkina Faso Sida’s contribution will significantly build in-country capacity to proactively respond to the pandemic May 21, 2020 – SEOUL, South Korea – The […]
INOVIO and GeneOne Life Science Report Positive Phase 1/2a Clinical Data With DNA Vaccine INO-4700 for MERS Coronavirus at the American Society of Gene & Cell Therapy (ASGCT) Conference
INO-4700 (GLS-5300) DNA vaccine demonstrates 100% binding and 92% neutralizing antibody responses against MERS-CoV INO-4800 DNA vaccine for COVID-19 currently in Phase 1 trial utilizes identical strategy targeting Spike protein and CELLECTRA intradermal delivery PLYMOUTH MEETING, Pa. and SEOUL, KOREA – April 28, 2020 […]
LINE and International Vaccine Institute Team Up to Promote the Importance of Vaccination
LINE to offer an Official Account to IVI to promote the importance of vaccines and vaccination, as well as feature educational contents starring IVI Goodwill Ambassador Henry Lau LINE FRIENDS’ BT21 character IVI sticker set to be launched in the first half of the year […]
IVI, INOVIO, and KNIH to partner with CEPI in a Phase I/II clinical trial of INOVIO’s COVID-19 DNA vaccine in South Korea
The Coalition for Epidemic Preparedness Innovations (CEPI) grants funding $6.9 million to INOVIO and IVI to conduct clinical testing in Korea for INOVIO’s COVID-19 vaccine candidate based on their well-established DNA platform technology Korea National Institute of Health (KNIH) to support IVI’s testing efforts […]
International Vaccine Institute and Technical University of Denmark to strengthen external quality assurance in the face of rising antimicrobial resistance in Asia
The Asia Pacific region is vulnerable to the emergence and spread of AMR but there is little high-quality data available on the extent of its impact Quality-assured data is essential for building tailored strategies for preventing the spread of drug-resistant infections IVI is part of […]
IVI and Sweden renew partnership to accelerate vaccines for global public health
January 13, 2020 – SEOUL, South Korea – The International Vaccine Institute (IVI), a Seoul, Korea-based international organization, announced today that the Swedish International Development Cooperation Agency (Sida) will continue to support IVI’s mission to accelerate vaccine research and development for global health […]
IVI to lead critical standard reagents availability for oral cholera vaccine manufacturing to ensure their uniform efficacy and help meet global demand
January 9, 2020 – SEOUL, South Korea – The International Vaccine Institute (IVI) received a $1.4 million grant from the Bill & Melinda Gates Foundation to ensure critical standards and reagents are available to low-cost oral cholera vaccine (OCV) manufacturers in the global health […]
IVI appoints three new members to its Board of Trustees
December 18, 2019 – SEOUL, South Korea – The International Vaccine Institute (IVI), a Seoul, Korea-based international organization dedicated to accelerating vaccines for global health, announced today that Professor Gordon Dougan, Dr. Melanie Saville, and Dr. Jean-Marie Okwo-Bele will join its Board of Trustees (BOT) […]
IVI to Accelerate Efforts in iNTS Vaccine Development
An award from the Wellcome Trust’s Affordable Innovations for Global Health Flagship will support IVI’s development and selection of potential vaccine candidates There is currently no vaccine available to protect against invasive non-typhoidal salmonella (iNTS) November 13, 2019 – SEOUL, South Korea – The […]
IVI acquires GCLP certification from Korean Ministry of Food and Drug Safety
Institute expected to increase contributions to vaccine development activities worldwide November 1, 2019 – SEOUL, South Korea – The International Vaccine Institute (IVI), an international organization for vaccine development and delivery headquartered in Seoul, has been granted Good Clinical Laboratory Practice (GCLP) certification by […]
IVI Awarded $4.5 Million Grant to Increase Accessibility of Oral Cholera Vaccine
October 28, 2019 – SEOUL, South Korea – The International Vaccine Institute (IVI) has received a $4.5 million grant from the Bill & Melinda Gates Foundation to simplify the current oral cholera vaccine (OCV) and further increase its accessibility. To date, the Gates Foundation has […]
IVI Discusses Expanding Support for Global Collaboration on Vaccine R&D and Delivery with Top Foreign Diplomats in Seoul
IVI State Forum 2019 was attended by 8th UN Secretary-General Ban Ki-moon, as well as 30 distinguished guests, including 17 ambassadors and top diplomats in Seoul, and senior Korean government officials. The IVI State Forum 2019 was attended by ambassadors and senior diplomats from […]
IVI-led Consortium Awarded Fleming Fund Regional Grant to Improve Data Sharing in Fight against Antimicrobial Resistance
September 24, 2019, SEOUL, Korea – The Chief Medical Officer for England, Dame Sally Davies, announced today during the 74th UN General Assembly that a consortium led by the International Vaccine Institute (IVI) has been awarded a £2.7m Fleming Fund Regional Grant to improve data […]
IVI to help developing countries tackle the spread of infectious diseases by building capacity through vaccinology training in Seoul
19th IVI International Vaccinology Course brings together 175 trainees and faculty members from 49 countries The annual course has trained over 1,544 healthcare professionals in vaccine development, evaluation, production and policy to help increase developing nations’ capacity in vaccine research and immunization. September […]
IVI acquires supplemental funding for Severe Typhoid in Africa Program (SETA Plus)
August 28, 2019 – SEOUL, South Korea – The International Vaccine Institute (IVI) has received additional funds of $4.33 million from the Bill & Melinda Gates Foundation to expand Severe Typhoid in Africa (SETA) Program activities in close alignment with the “Effect of a […]
IVI Director General Dr. Jerome Kim to join the Human Vaccines Project’s Efforts as One of Four Distinguished Global Leaders to Decode the Human Immune System
August 16, 2019 – SEOUL, South Korea – Dr. Jerome Kim, Director General of the International Vaccine Institute (IVI), joins the Human Vaccines Project as one of four distinguished global leaders to help push forward its effort to solve one the greatest public health challenges […]
Prof. Young Chul Sung, biotech leader, personally donates US$8.4 million to IVI
Fund to support IVI’s projects to vaccinate children in developing countries throughout Asia-Pacific and Africa, to develop vaccines against emerging viruses Prof. Park Sang-chul, President of the Korea Support Committee for IVI, Prof. Sung Young-chul with his donation pledge and IVI Director General Dr. […]
New push to develop world’s first vaccine against the deadly Strep A bacteria killing hundreds of thousand
Streptococcus pyogenes bacteria, which causes scarlet fever and other infections 30 May 2019, SEOUL, South Korea – The International Vaccine Institute (IVI) and Australia’s Murdoch Children’s Research Institute (MCRI) will coordinate a global push to free the world of Group A Streptococcus (Strep A), […]
IVI-led CAPTURA consortium wins Fleming Fund award for work on antimicrobial resistance (AMR) data across Asia
SEOUL, South Korea — An international consortium, led by the International Vaccine Institute (IVI), has received funding from the UK’s Fleming Fund Regional Grants to conduct the Capturing Data on Antimicrobial Resistance (AMR) Patterns and Trends in Use in Regions of Asia (CAPTURA) project. The […]
IVI receives $15.7 million grant to conduct phase III trials of new Vi-DT typhoid conjugate vaccine manufactured by SK bioscience of Korea
– Grant from the Bill & Melinda Gates Foundation will support additional supply of novel TCV at affordable cost for public sector markets SEOUL, Korea –The International Vaccine Institute (IVI) announces receipt of a $15.7 million-dollar grant from the Bill & Melinda Gates Foundation […]
‘IVI Forum for Enhancement of Cooperation’ hosted jointly by the Korean Parliamentary Forum for Global Health, and KCDC under MOHW
– Event organized by IVI to discuss ways of cooperation to help increase Korea’s vaccine self-sufficiency, and accelerate globalization of the Korean vaccine industry Key participants pose for a commemorative photo at the ‘IVI Forum for Enhancement of Cooperation’ at the National Assembly of the […]
Vaccine study confirms sensitivity of cholera test
Recently, the sensitivity of fecal microbiological cultures for detecting cholera has come under question. Researchers reporting in PLOS Neglected Tropical Diseases investigated this claim using a ‘vaccine probe’ analysis of a completed cholera vaccine cluster randomized trial to support the sensitivity of conventional microbiological culture […]
Korean Pediatric Association, IVI, KSC exchange MOU
KPA, as group of experts in pediatric infectious diseases, to collaborate in academic research and projects on infectious disease prevention To support vaccination of children in developing countries through the ‘One for Three’ campaign, in partnership with IVI The International Vaccine Institute (IVI), […]
IVI receives $3.2 million grant to support TCV effectiveness studies in West Africa
SEOUL, Korea — The International Vaccine Institute (IVI) has been awarded a $3,238,974 grant from the Bill & Melinda Gates Foundation to provide technical assistance support for studies to measure the effectiveness of typhoid conjugate vaccine (TCV) in West Africa. This grant comes after […]
CEPI partners with IVI to accelerate development of vaccines against emerging global health threats
– Bilateral ‘Master Implementing Partner Services Agreement’ expected to enable IVI to provide technical capabilities to CEPI Oslo, Norway; Seoul, Republic of Korea 11 February 2019—The Coalition for Epidemic Preparedness Innovations (CEPI) and the Republic of Korea-based International Vaccine Institute (IVI), an international organisation […]
Henry Lau joins IVI as Goodwill Ambassador
– Singer-songwriter, Henry to team up with the International Vaccine Institute (IVI), founded by the United Nations Development Programme to develop and deliver new vaccines for global health – IVI Director General Dr. Jerome Kim: “IVI is privileged to have Henry on board, and grateful […]
Nature Communications study addresses MDR typhoid
New study in Nature Communications reveals high burden of multi-drug resistant (MDR) typhoid fever in sub-Saharan Africa associated with two predominant genotypes – IVI epidemiologist Se Eun Park and collaborating scientists report data from Typhoid Fever Surveillance in Africa (TSAP) program and other sources […]
THSTI, IVI held joint symposium on Nov. 22
First cooperative program between International Vaccine Institute and Translational Health Science and Technology Institute expected to help accelerate cooperation between India and IVI IVI and the Translational Health Science and Technology Institute (THSTI) of India held a joint symposium at IVI headquarters on November 22. […]
‘The Euvichol story’ published in Vaccine
“The Euvichol Story – Development and licensure of a safe, effective and affordable oral cholera vaccine through global public private partnerships” spearheaded by the International Vaccine Institute (IVI), has been published in Vaccine by Elsevier. The Euvichol Story published on October 9, 2018 in the […]
IVI accelerates development of MERS vaccine
– Institute jointly conducts phase 1/2a clinical trial of GeneOne Life Science’s GLS-5300 DNA MERS vaccine – IVI in search of additional promising candidate vaccines among developers worldwide, with support from Samsung Life Public Welfare Foundation The International Vaccine Institute (IVI) said on September […]
IVI’s 18th Vaccinology Course gathers 120 international participants from 19 countries in Seoul September 3-7
Annual Course to update on the recent development in vaccinology, including basics of immunology, epidemiology, vaccine design and development Featured experts include: Stanley Plotkin (University of Pennsylvania), Dr. Barney Graham (US NIH), Dr. Randy Schoepp (USAMRID), Dr. Andy Pollard (University of Oxford), and Dr. Eric […]
KOICA, IVI team up with Mozambican MoH and INS, partners to vaccinate 190,000 people against cholera
– Joint initiative aims to prevent and control cholera, and secure sustainable cholera and diarrheal disease surveillance – WASH campaign to promote access to clean water and behavioral change for appropriate sanitation and hygiene practice – “KOICA is joining the program to contribute to ‘Ending […]
IVI acquires $5 million grant to support process development, scale up of typhoid conjugate vaccine with SK Bioscience
– Grant from the Bill & Melinda Gates Foundation to allow SK Bioscience Co. Ltd, IVI to accelerate late-stage development of new vaccine necessary to achieve WHO prequalification The International Vaccine Institute (IVI), an international nonprofit organization devoted to providing vaccines critical to global public […]
×
Evolutionary mechanism of cholera bacterium identified
|
||||||||
|
||||||||
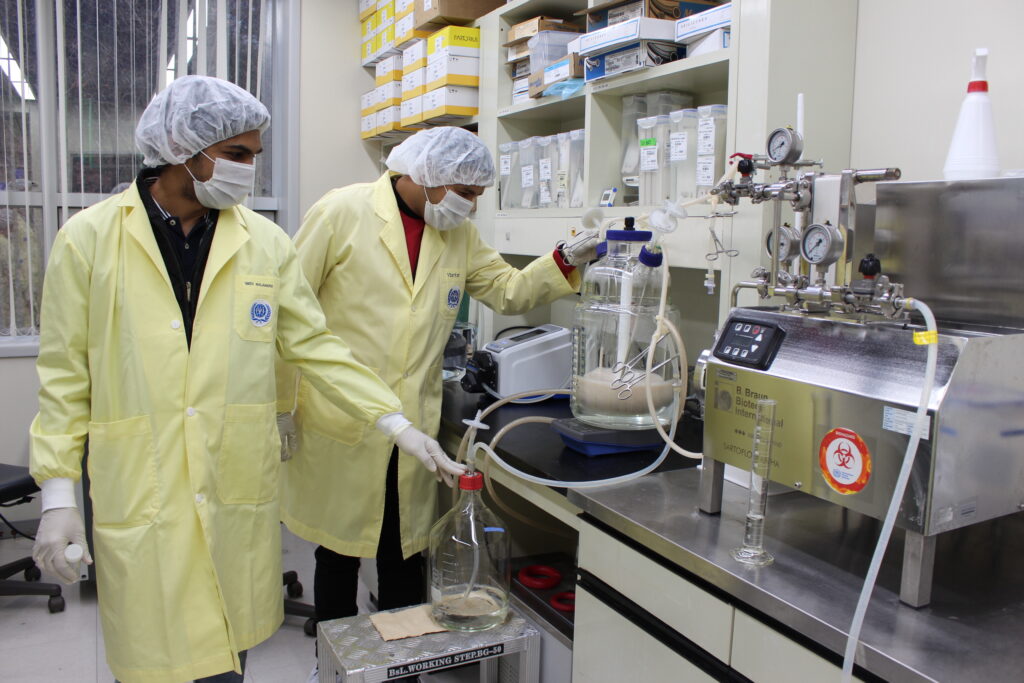
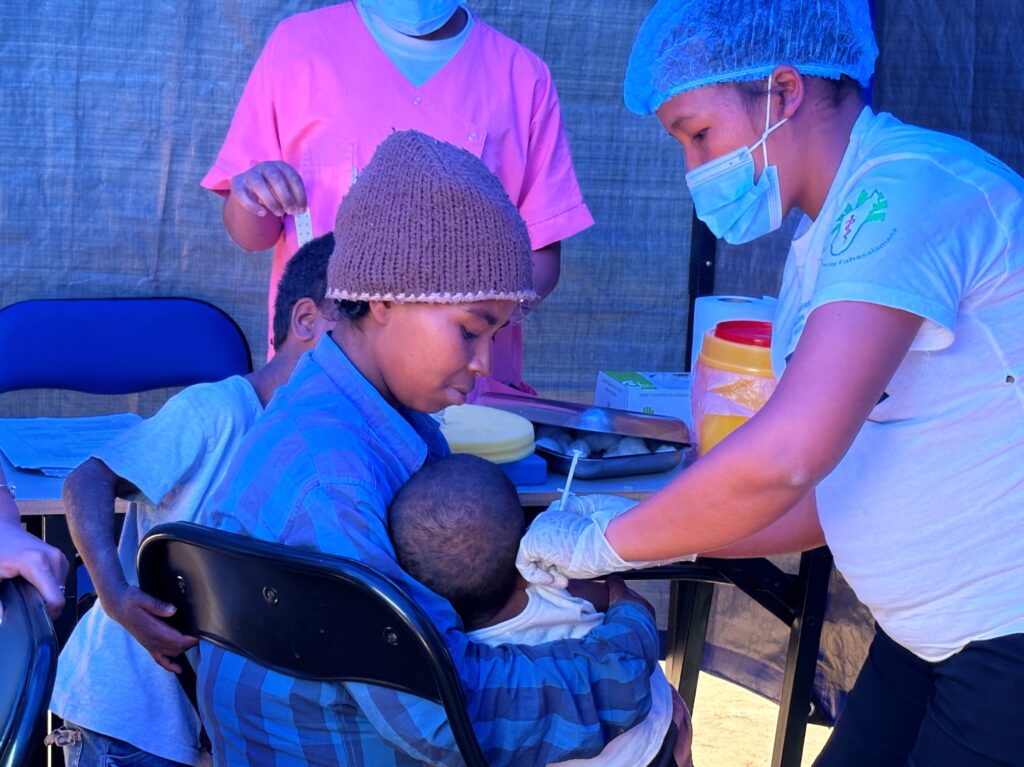
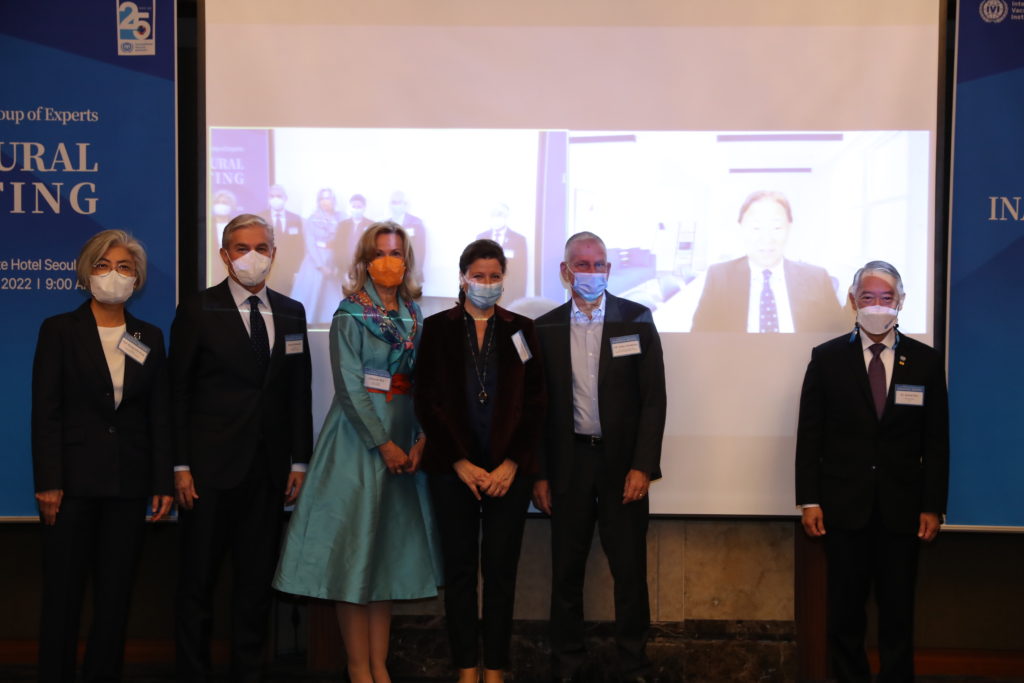
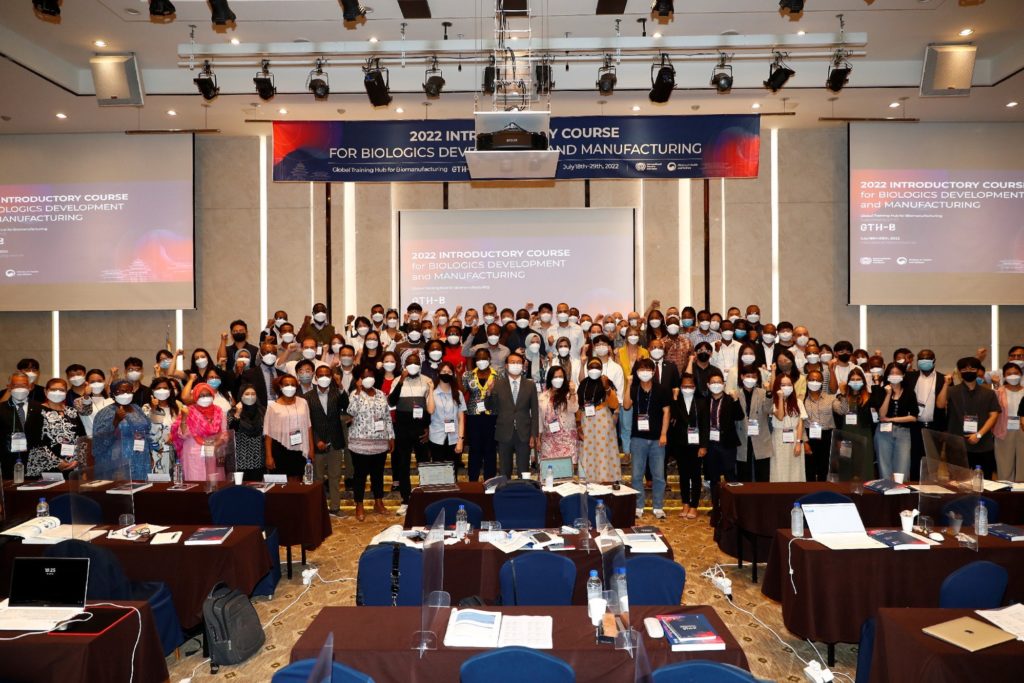
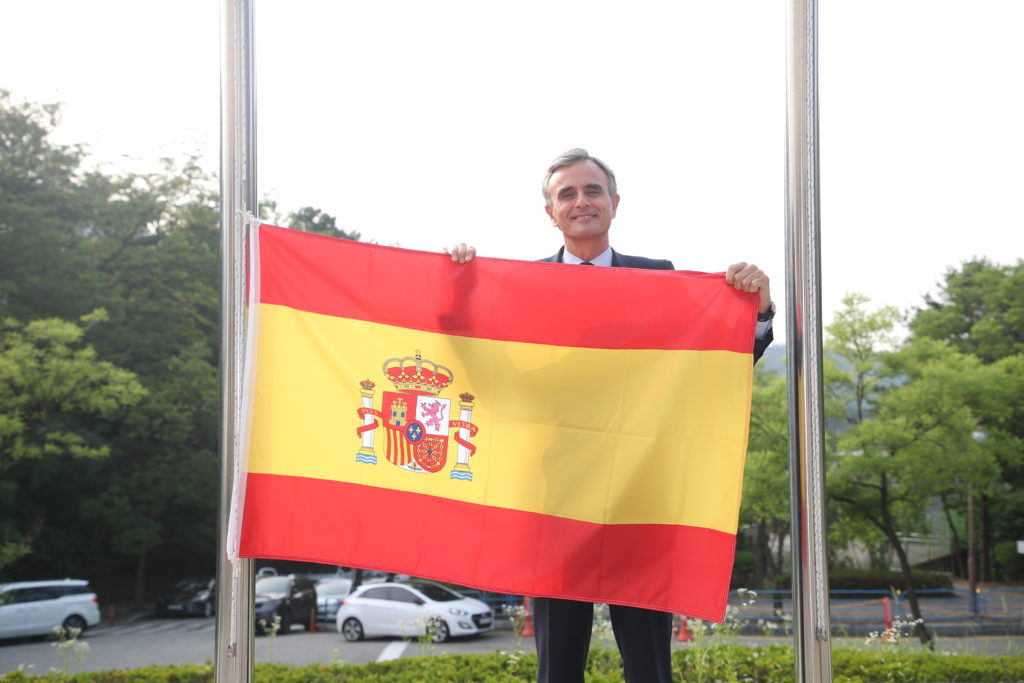
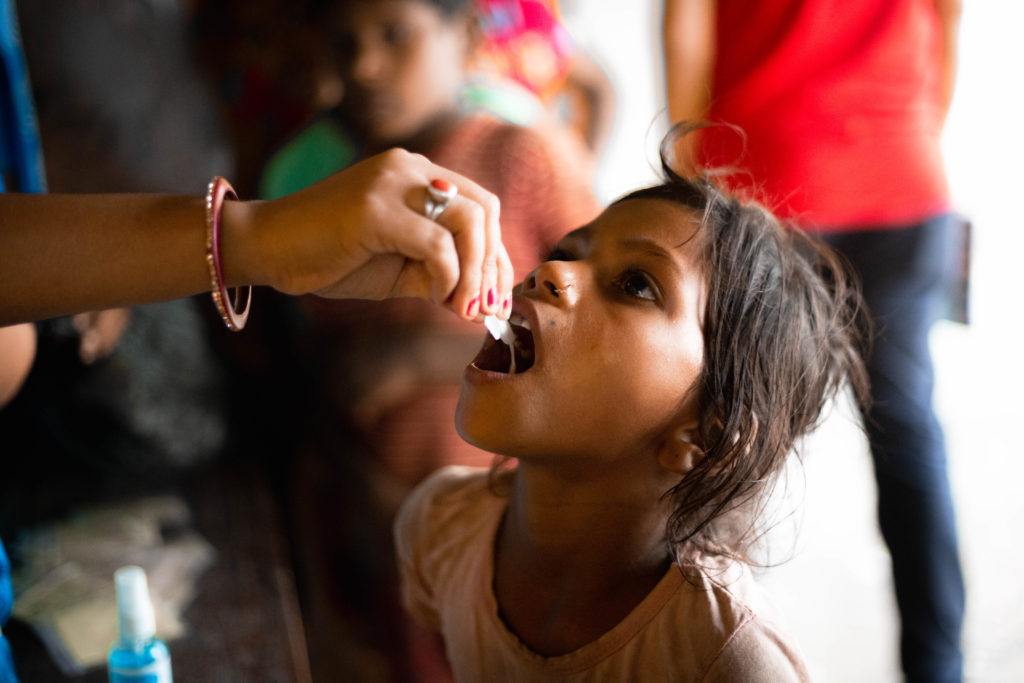
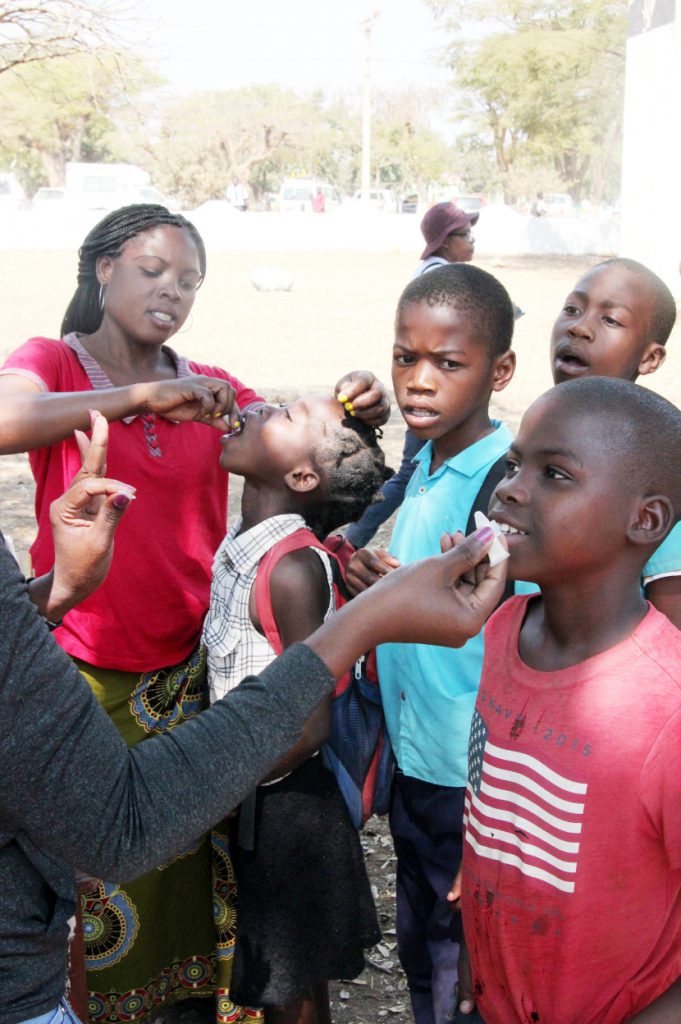
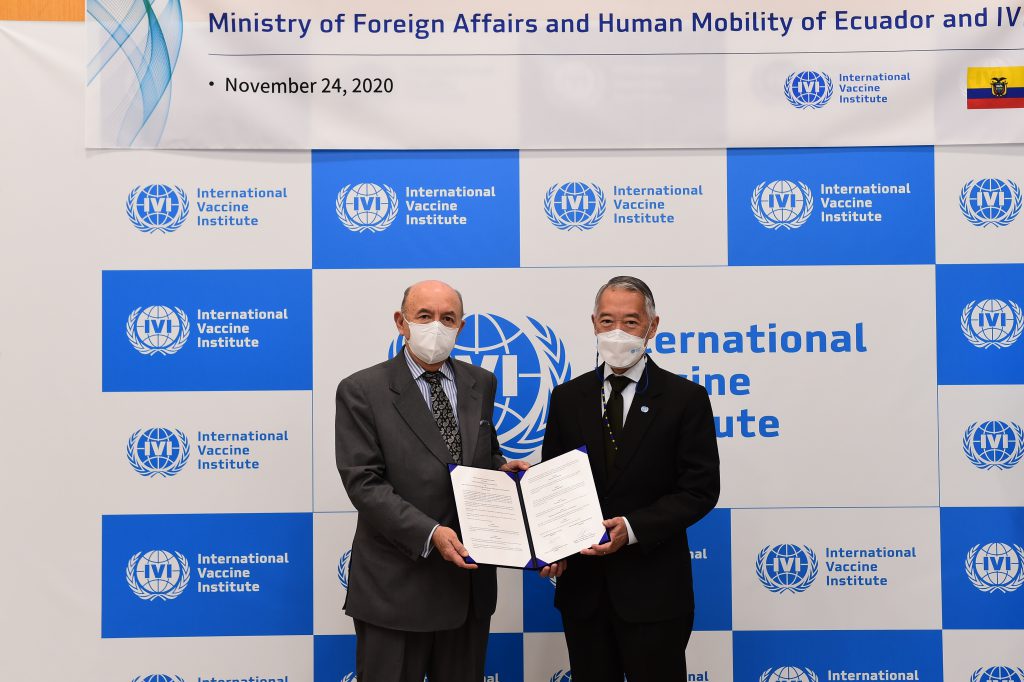















































































































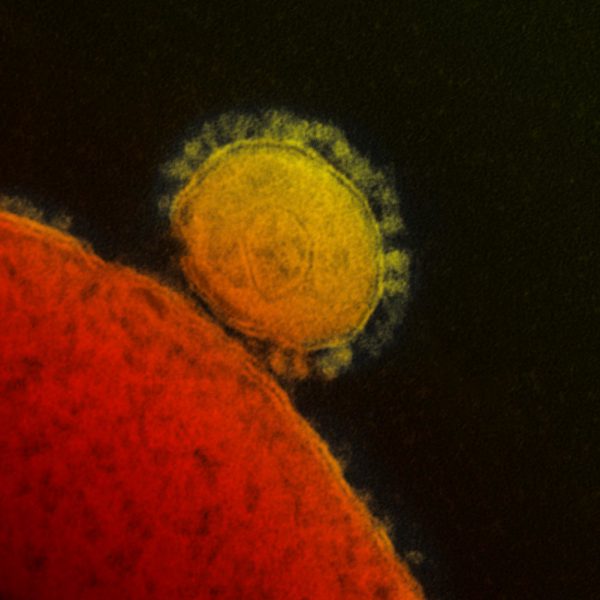









































































































































 An international research team comprising of scientists from Seoul National University (SNU), the International Vaccine Institute (IVI) in Korea, and U.S. research institutions elucidated the basis for the emergence of mutated strains of Vibrio cholerae, the causative agent of cholera, and identified an evolutionary mechanism of pathogenic bacteria by analyzing genomic sequences of 23 V. cholerae strains.
An international research team comprising of scientists from Seoul National University (SNU), the International Vaccine Institute (IVI) in Korea, and U.S. research institutions elucidated the basis for the emergence of mutated strains of Vibrio cholerae, the causative agent of cholera, and identified an evolutionary mechanism of pathogenic bacteria by analyzing genomic sequences of 23 V. cholerae strains.